Tracking the Evolution of Cancer, Cell by Cell
In August, when a group of physicians writing in the Journal of the American Medical Association proposed that some precancerous conditions should no longer be labeled as cancer, the recommendation was supported by studies showing that advances in the detection of these conditions haven’t reduced the incidence of invasive cancers. Indeed, precancerous conditions such as ductal carcinoma in situ — abnormal cells in part of the breast that have not spread — can progress to cancer but often don’t, raising questions about how aggressively to treat them. The larger issue is that physicians don’t yet have a reliable way to distinguish precancerous cases that will remain harmless from those that will become invasive.
This challenge, and many others that stymy both physicians treating cancer and scientists trying to understand it, can be traced to cancer’s inherent heterogeneity. Cancer cells possess unstable genomes — meaning their chromosomes can gain and lose pieces with disturbing speed — and they grow rapidly, creating a genetically diverse population. As a result, different cells within the same tumor can look quite different, with different numbers of chromosomes and unique mutations, for example. Only a subset of cells within the tumor may carry mutations that allow them to grow unchecked, to spread to a new environment or to evade a specific drug. But because most cancer research averages the differences from many cells, these rare mutations, which lie at the heart of cancer’s deadliness, may get lost in the balance.
That may soon change. New technology is allowing researchers to more precisely track the genetics of individual cancer cells. Whereas earlier studies could look only at rough measures, such as the number of chromosomes in a cell, scientists can now record the number of copies of multiple genes in each cell and in some cases read the entire genome, which gives a more precise picture of the genetic diversity among single cells. Recent studies using these techniques to compare cells from different parts of the same tumor and over time have revealed their extreme diversity, providing an intimate portrait of the mutations that give the disease its power. Knowing more about the rate of mutations and the drivers of genomic instability may help answer fundamental questions about the origins of cancer, which could in turn improve treatment options.
A study published last year analyzing 100 or more individual cancer cells from each of 13 women who had both pre-invasive ductal carcinoma and invasive breast cancer found that key cancer genes varied in number from cell to cell. Even the pre-invasive cells were highly genetically unstable. That supports the idea that this pre-invasive condition should also be treated, because the cells are already genetically diverse and may therefore harbor the ability to spread. (However, scientists need to confirm that pre-invasive ductal carcinoma in women who don’t have invasive cancer is similarly genetically unstable.)
Studies in population biology and ecology suggest that a more genetically diverse population is more robust, meaning more likely to survive in the face of environmental challenges. Nicholas Navin, a cancer geneticist at the University of Texas’ MD Anderson Cancer Center in Houston, theorizes this will also be true for cancer. “Diversity allows tumors to deal with selective pressures,” he said. In a genetically diverse tumor, millions of cells will die when exposed to a drug, but there’s a good chance that a few cells will be genetically empowered to survive. These cells will experience a ‘population bottleneck’ — when most are killed off — “but then continue to survive and repopulate the tumor mass over time,” Navin said.
Diverse and Deadly
Alejandro Schäffer’s mother and first wife died of esophageal and stomach cancer, respectively. Then, two years ago, his father was successfully treated for esophageal cancer with a novel drug. “I have seen firsthand that recent developments in cancer research can cure some patients, but there is a long way to go,” said Schäffer, a computational biologist who began studying the disease soon after he joined the National Institutes of Health nearly 20 years ago.
In 2008, while visiting his recently widowed father in Pittsburgh, Schäffer met Russell Schwartz, a computer scientist at Carnegie Mellon University. Like Schäffer, Schwartz was working on an evolutionary approach to cancer. According to this view, cancer cells evolve and adapt to their environment, just as species of salamanders might adapt to a changing climate or spread to inhabit new territory. This evolutionary perspective, laid out nearly 40 years ago, spawned decades of theoretical research and shaped scientists’ views of how cancer originates and spreads. But little direct DNA evidence supported these notions. Now, equipped with new techniques to analyze the molecular signatures of individual cancer cells, paired with tools from evolutionary biology, scientists are beginning to track cancer’s evolution at a level of detail that could help them pinpoint the mechanisms behind its wily adaptation and unfettered growth.
“It is becoming increasingly appreciated that tumor progression is an evolutionary process of genomic changes,” Schäffer said by e-mail. “Therefore, techniques developed to understand evolution of species can and should also be applied to understand tumor progression.”
For both salamanders and cancers cells, reproduction provides a particularly risky time for the genome. The chromosomes must replicate before lining up like soldiers, splitting into regiments and marching to new bunkers in daughter cells. Each stage of the process presents an opportunity for error — mistakes can occur as DNA is copied, and chromosomes can misalign, break apart or wind up in the wrong cell. From these genetic gaffes, cancer is born.
In the evolutionary approach to cancer, each cell type is treated as an individual species and plotted onto a phylogenetic tree, a structure that maps out the evolutionary relationship between organisms. The tree structure makes it possible to calculate, for example, which genes are most likely to be copied or deleted when the cell divides and which parts of the tree are the best competitors in terms of growth and survival, Schäffer wrote.
This approach allows scientists to look at more than a single biomarker, such as a gene or protein, or at the diversity of a cancer at one point in time, but at the dynamics of the tumor itself, which may better predict its behavior. “The issue is to understand how important dynamics are for cancer progression,” said Thomas Ried, an oncologist at the National Cancer Institute in Bethesda, Md., who works with Schäffer and Schwartz.
In a paper published in the journal Bioinformatics in July, the researchers used this approach to try to answer one of the most-challenging questions facing oncologists: What makes some cancers spread? “Evolutionary trees become a way of thinking about the evolution process in tumors,” Schwartz said.
Animal Models
To get an accurate picture of the diversity of cancer, scientists would like to be able to analyze individual cells both at different places in a tumor and at different times as a cancer develops. This is difficult to do in people with solid tumors because it requires invasive biopsies. But scientists are now beginning to carry out such experiments in animal models of cancers.
They built phylogenetic trees using single-cell data from women with cervical cancer, half of whose cancers had spread. They then used machine-learning methods, computer algorithms designed to look for predictive features in complex data, to compare the structure of the trees. A narrow tree with fewer branches, for example, has less genetic variability than a bushy one. “We can ask what’s in common among trees or what separates tumors that do progress and those that don’t,” Schwartz said.
Overall, the tree approach was better able to predict which of the primary tumors later metastasized than standard biomarkers, such as the number of copies of an individual gene. Scientists haven’t yet pinpointed the key predictive feature within the phylogenetic trees, but Schwartz and his colleagues found that metastatic cancers have narrower trees, which may reflect the selective pressure imposed on cells that are spreading across the body. “It requires a more specific set of mutations to enable a tumor to migrate to and survive in a distant site versus its tissue of origin,” he said.
Predicting Harm
With these preliminary findings, scientists are beginning to test the single-cell approach on other data sets, but they haven’t yet used it to predict that a cancer will spread before it actually has. Nevertheless, they highlight the potential for this approach to answer both fundamental and clinically relevant questions about the disease.
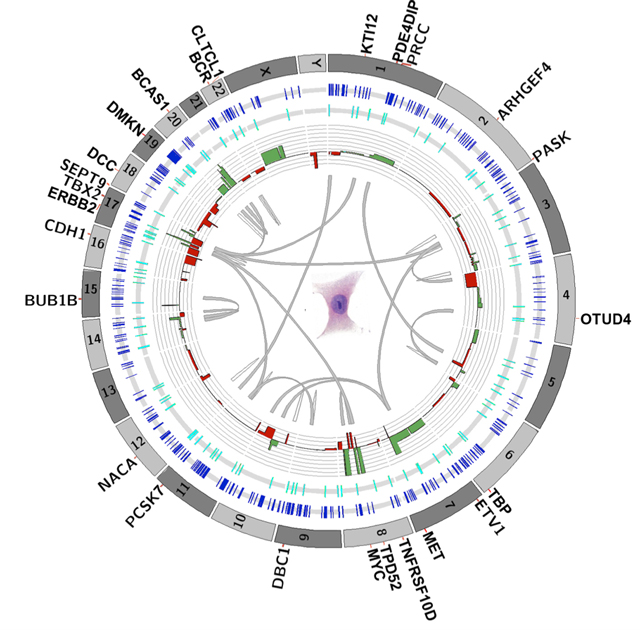
Schwartz’s study aimed to predict cancer severity by looking at a handful of genes within each cell. Others are now starting to examine the entire genome of single cells, providing a more comprehensive evolutionary picture. In 2011, Navin, then a postdoctoral researcher at Cold Spring Harbor Laboratory in Long Island, and collaborators published the first analysis of single-cell genomes in cancer, sequencing the full genome of 100 cells in each of two cases of breast cancer. The results challenged the predominant dogma, that cancer cells accumulate genetic changes gradually.
“Instead of acquiring mutations over 10 to 20 years, our data suggested evolutionary bursts with lots of genomic mutations,” Navin said. That burst might be triggered by environmental exposure to X-rays or a mutation in a gene that helps to repair DNA, pushing a cell over the edge and into the realm of genomic instability. “After a punctuated burst of genome rearrangements, a cancer cell might already have the proper mutations to become invasive, metastatic and ultimately kill the patient,” he said. “Once you acquire these chromosome aberrations, you are at a point of no return.”
Mutations Large and Small
Studies of single-cell sequencing suggest that large genetic changes, which can include multiple genes, accumulate in cancer cells in bursts. But preliminary research by Navin and collaborators suggests that point mutations — changes to individual DNA letters — accumulate more slowly.
Navin said the two models are similar to those proposed for speciation: Stephen Jay Gould’s idea of a punctuated model for how species evolve versus Darwin’s original model of gradual changes. The two models aren’t mutually exclusive, however, and the two processes run according to different molecular clocks.
Moving forward, Navin and other researchers plan to put cancer’s reputation for accelerated mutations to the test. While the working assumption has been that cancer cells acquire more mutations per division than healthy cells, it’s also possible that they simply divide more rapidly, racking up a greater record of mutations in the same amount of time. “That’s something that’s very contentious,” Navin said. “With single-cell sequencing tools, we can really dissect these mutation rates, which hasn’t been possible before.” The answer could have implications for treatment — cells that divide more rapidly, for example, would be more vulnerable to drugs that effect replication or the cell cycle.
On the clinical side, researchers plan to examine why some cancers bounce back months or years after treatment. In the past decade, targeted cancer therapies, which are designed to block specific molecular changes in cancers, have made major strides in treating the disease. But their success is typically short-lived. People treated with these drugs often relapse, suggesting that their tumors have become resistant. “I think it is becoming apparent that intra-tumor heterogeneity is one of several major impediments to personalized cancer treatment by single ‘magic bullet’ chemotherapies,” Schaffer said.
For example, some tumors may harbor rare cells capable of resisting certain drugs. Analyzing single cells may ultimately help scientists and doctors better predict when a cocktail of drugs targeted at different molecular mechanisms, similar to the approach in HIV treatments, is warranted. “Understanding tumor heterogeneity will hopefully help end the problem of disease recurrence,” Ried said.
This article was reprinted on ScientificAmerican.com.