Scientists Map 5,000 New Ocean Viruses
In March 2011, the Tara, a 36-meter schooner, sailed from Chile to Easter Island — a three-week leg of a five-year global scientific expedition. All but one of the seven scientists aboard the ship spent much of their time on the sun-drenched deck hauling up wondrous creatures such as luminous blue jellyfish and insects known as sea-skaters, which spend their entire lives skimming the surface of the ocean far from land.
At the stern of the Tara, a shipping container was bolted to the deck, with a door and a tiny window cut through the metal walls. One of the scientists, Melissa Duhaime, spent most of the voyage inside the dark, tiny cell, where she fought off an endless bout of seasickness.
“People would come in to see what I was doing and leave pretty quickly,” Duhaime said.
Inside her cell, Duhaime sat next to a hose as wide as an outstretched hand. A pump drew water through the hose from several meters below the boat and then pushed it through a series of filters. Each filter was finer than the last, blocking smaller and smaller life forms. The setup stopped animals first, then zooplankton and algae. The last filter in the hose, with pores just 220 nanometers wide, was fine enough to block bacteria. Scrubbed of all these living things, the water finally flowed into three 30-liter vats.
To the untrained eye, these vats might seem to be full of sterile water. But they were seething with ocean life — or life-like things, at the very least. The three vats held up to 1 trillion viruses.
The ocean contains many mysteries, but none so great as its viruses. Scientists estimate that there are 1,000,000,000,000,000,000,000,000,000,000 virus particles in all the world’s seas. They outnumber all cellular life forms by roughly a factor of 10. Scientists have been dimly aware of the staggering scale of the ocean’s virosphere since the late 1980s, but many of the simplest questions about it remained open for years. Scientists couldn’t even say how many species of viruses there were in the oceans. It’s as if zoologists were dimly aware that many places on Earth were home to things called mammals, but their knowledge was based only on a few squirrels in a cage.
Duhaime and her colleagues joined the Tara Oceans Expedition to change that, by collecting ocean viruses on a scale never attempted before. As they report in the May 22 issue of Science, they gathered enough samples to confidently estimate the total number of distinct populations of viruses in the sunlit upper reaches of the ocean. Out of the 5,476 populations they identified, only 39 were previously known to science.
The researchers have gone on to study where these species live and how they affect the ocean’s ecosystems. For years, scientists have suspected that viruses alter the very chemistry of the world’s oceans and may even influence the planet’s climate. Now, the data from the Tara are going to give researchers a much better handle on the full power of the ocean’s viruses.
To Duhaime, who is now at the University of Michigan, getting a glimpse of the ocean’s virosphere made three seasick weeks in a darkened metal hut worth the discomfort. “You’re in this moment in nature where all these important cycles are going on all around you in the ocean. You’re just trying to take a snapshot of that,” she said. “That never gets old.”
An Explosion of Viruses
Scientists first discovered viruses in sickly tobacco plants in the late 1800s. Yet for nearly a century, marine biologists assumed that the oceans were virtually virus-free. When they looked at seawater under microscopes, they simply didn’t see any viruses, and so they concluded that the oceans were too harsh for viruses to survive in great numbers.
In the 1980s, a biologist named Lita Proctor decided to take a more careful, systematic look. Her surveys of water from places like the Caribbean and the Sargasso Sea revealed a surprising abundance of viruses. Other researchers went on to confirm that the oceans are indeed a viral soup.
It became clear that viruses are an important part of the ocean’s ecology, but scientists struggled to study them. Simply staring at a virus through a microscope doesn’t tell you all that much about it. Two viruses that look nearly identical may infect completely different hosts. Since scientists couldn’t tell which hosts the viruses needed in order to replicate, they struggled to rear them in the lab.
Then in the 1990s, a new way to survey life emerged. Scientists would add chemicals to a sample of seawater (or soil or lake mud or some other material) that would rip apart all the proteins and membranes it contained. Out of that detritus, the scientists could extract all the DNA from the sample in a jumble of fragments. The researchers then sequenced the fragments and pieced them together into larger DNA segments. Finally, they could compare these genetic sequences to those of known species — finding either an exact match, or a sequence from a closely related species.
This method, known as metagenomics, quickly gave scientists a wave of new discoveries about bacteria and other microbes. It transformed the study of human health by allowing scientists to catalog the thousands of species of microbes that live in and on the human body, which had previously been unknown because they couldn’t be grown outside of our inner jungles.
But ocean viruses don’t surrender their secrets so easily to metagenomics. All cellular species, from E. coli to fin whales, have a core set of genes in common. Viruses, on the other hand, have no such universal set of genes. When scientists gather genes from a virus that’s new to science, they often find that almost none of its genes bear any resemblance to any previously discovered viral gene. In addition, viruses often pick up new genes, either from other species of viruses, or from their hosts. When scientists isolate one piece of genetic material from an unknown virus, it can be difficult to determine where it came from.
In recent years, scientists have managed to bring some order to this chaos. Matthew Sullivan, an environmental virologist at the University of Arizona, and his colleagues have developed a method to identify new species of viruses. They start with viruses that share some of the same genes. Then they measure how similar their DNA is. When they plot those measurements on a graph, the viruses do not spread out into a hazy, inscrutable cloud. Instead, they huddle in tight clumps. These clumps, Sullivan and his colleagues argue, represent distinct species of viruses.
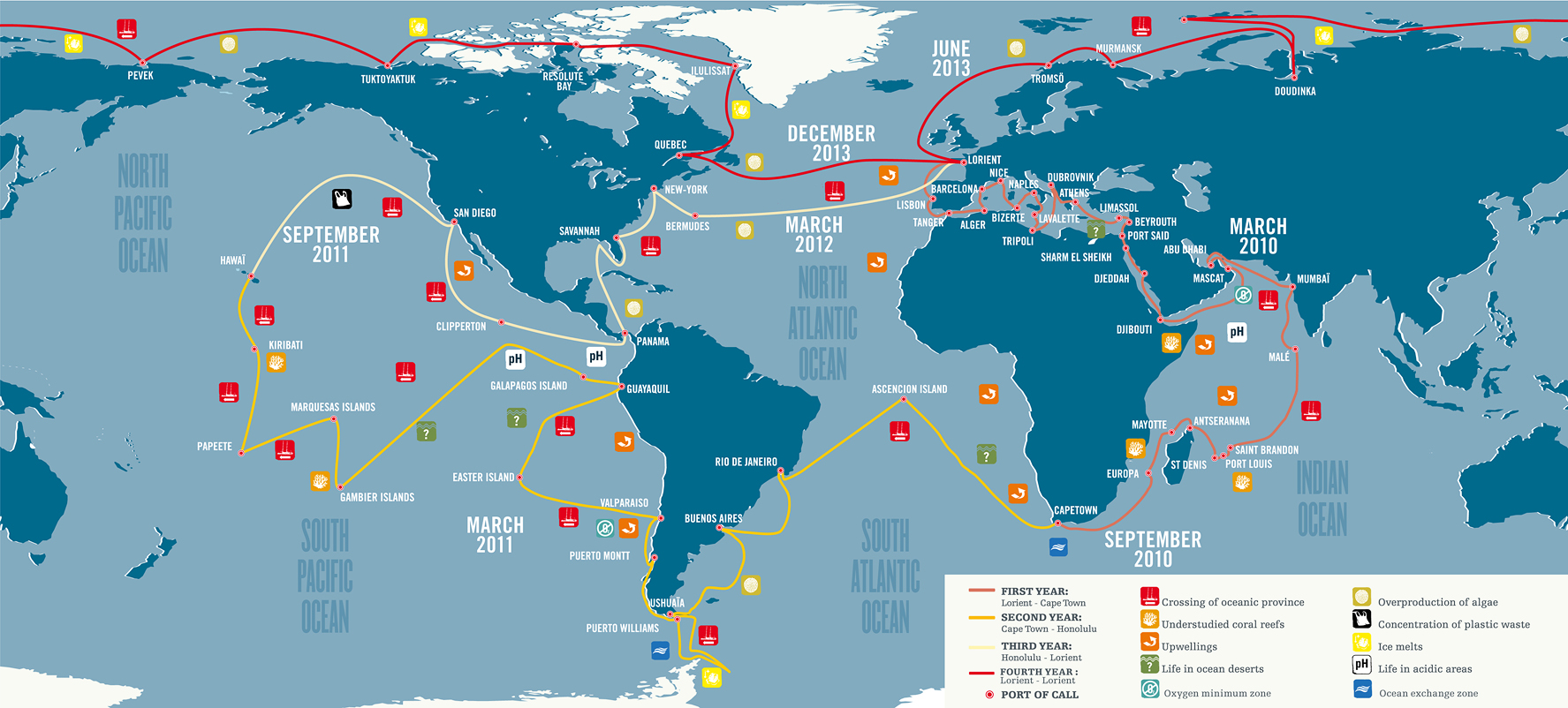
Sullivan joined the leadership of the Tara Oceans project to put these new methods to work on a global scale. His team of researchers filtered seawater in every ocean on Earth, along with the Red Sea, the Mediterranean and the Adriatic, visiting 43 sites in all. They processed the water with iron chloride to make the viruses stick together in particles, filtered the water again, and shipped the virus-clogged filters to labs around the world.
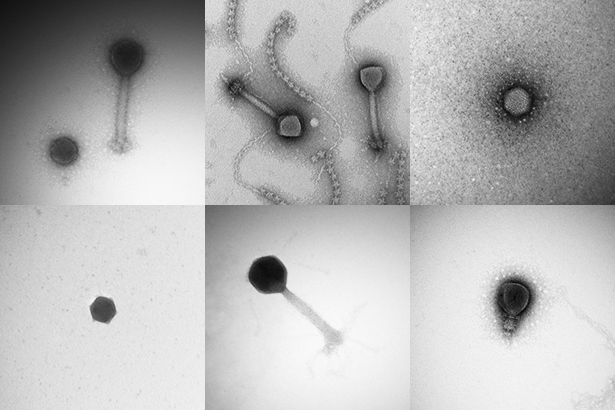
Sullivan and his colleagues examined the viruses under microscopes and sequenced their genes. All told, they extracted 2.16 billion pieces of DNA, each piece containing 101 base pairs on average. The scientists found that many of the pieces overlapped with each other. In jigsaw-puzzle fashion, they assembled long stretches of DNA and began to recognize the full sequence of virus genes.
As they compiled this genetic catalog, they began to sort it. Sullivan and his colleagues combined closely related genes into clusters. A pair of genes that encode for similar proteins would be placed into the same cluster. All told, they identified 1,323,921 clusters.
Now they were left with a crucial question: What fraction of all the virus genes in the world’s oceans had they collected? Had they just skimmed the surface, or did these 1.3 million clusters represent most of the viral genes in existence?
There’s a clever statistical trick scientists can use to answer a question like this. Sullivan and his colleagues began by randomly picking a gene from their catalog. Then they picked another one and noted whether the two genes belonged to the same cluster, or to two different ones. Then they picked out a third gene and compared it to the first two. At each step, they marked their progress by plotting a point on a graph, where every novel gene made the graph tick up.
At first the curve was steep, because each new gene typically didn’t belong to any cluster identified so far. But after a while, the curve flattened as the new genes fell into existing clusters. By the end of the process, it was rare for the scientists to pick a gene that was new. Repeating this exercise over and over again with randomly selected genes produced the same flattened curve.
This flattened curve told Sullivan and his colleagues something profound: that they have probably found almost all the virus genes in the upper oceans of the entire planet. There are not billions of additional genes lurking out there, waiting to be sucked into a hose.
“That’s nice,” Sullivan said, “because it’s a finite number we can work with.”
The scientists then used this catalog to figure out how many different kinds of viruses there are in the world’s oceans. (For now, Sullivan is careful to call these distinct kinds of viruses “populations,” but he’s confident that with further research, he’ll be able to show that these populations are, in fact, true species.) By comparing the genes in the clusters to one another, the scientists were able to identify 5,476 distinct populations.
With this single expedition, the scientists have dramatically increased our view of the ocean virosphere. When Sullivan and his colleagues tried to match their 5,476 populations to species that scientists have already documented, they succeeded just 39 times. In other words, 99 percent of the viruses they discovered were new to science.
The flattened curve of genes tells Sullivan and his colleagues that if they went out on a new expedition, using the same methods to identify viruses, they wouldn’t find many more new ones. But Sullivan is quick to point out that the total number of species of ocean viruses will turn out to be more than 5,476. In recent years, for example, scientists have found a number of so-called “giant viruses” that are as big as bacteria. The filters that Duhaime and others used on the Tara Oceans Survey prevented any giant viruses from getting into the vats they studied.
In addition, the scientists sequenced only viruses that use DNA to encode their genes. Some viruses, such as influenza and HIV, use RNA, a single-strand version of DNA, to encode their genes. By one estimate, as many as half of the viruses in the ocean are RNA viruses. What’s more, the Tara Oceans survey only took samples from the surface of the ocean. The deeper regions have viruses, too, as does the sediment at the bottom of the sea.
Nevertheless, the Tara Oceans survey leaves Sullivan confident that the total number of ocean viruses will only be in the tens of thousands.
“It’s a smaller number than I expected,” Sullivan said. Based on smaller studies, some scientists had speculated that there might be hundreds of thousands of species of ocean viruses and billions of viral genes. But the Tara Oceans survey suggests that’s not the case. “It is what it is,” Sullivan said.
What Viruses Do
With such a comprehensive picture of ocean viruses, Sullivan and his colleagues can start drawing some broad conclusions about them. Each virus population is more common in some areas than in others, for example. That’s likely due to the fact that their hosts thrive at certain temperatures or with certain levels of oxygen.
But Sullivan and his colleagues found viruses from most populations everywhere they looked. In other words, every part of the ocean is like a seed bank for viruses. As soon as the right host comes along, a relatively rare virus will infect it and replicate itself into a huge population.
Now that scientists have such a clear picture of the diversity of ocean viruses, they can hope to gain a better understanding of how these viruses are affecting the planet. Viruses kill vast numbers of hosts. Some researchers have estimated that they kill up to 40 percent of all bacteria in the ocean every day. Paradoxically, though, this daily massacre could actually increase the biomass of the oceans. Mathematical models of ocean ecosystems suggest that by killing so many microbes, viruses could release carbon and other organic nutrients back into the environment, providing an easy source of nutrients for other organisms. Carbon that isn’t consumed may drift down into the deep ocean, thereby causing vast supplies of carbon to fall to the seafloor rather than escape into the atmosphere.
Until now, scientists haven’t known enough about ocean viruses to create precise models of these effects. So they haven’t been able to say with confidence what the viruses are doing to our world. Duhaime said that the data coming from the Tara Oceans survey will go a long way toward pinning those models down. “We’re very far from doing that,” she said, “but the path is there.”
Despite all this new data, Sullivan still considers the ecology of ocean viruses to be in its infancy. “The virus stuff isn’t as mature as counting zebras,” he said. “That’s because it’s easier to observe zebras and define a zebra species than it is for viruses.”
But Sullivan and his colleagues have built a pipeline they can now use to pump huge numbers of additional viruses. Animal ecologists have a head start that stretches hundreds of years, but according to Sullivan, “we may quickly be ahead.”
Correction on May 22, 2015: Sullivan estimates that the total number of oceanic viruses will be in the tens of thousands; previous estimates had been in the hundreds of thousands.