Ancient DNA Yields Snapshots of Vanished Ecosystems

Under the right circumstances, loose DNA from expired animals, plants and microbes can often survive in nature for many thousands of years. Metagenomic techniques for studying this environmental DNA (eDNA) are helping researchers to glimpse microcosms of vanished ecosystems in bits of ice, sediment and soil.
Ryan Garcia for Quanta Magazine
Introduction
Somewhere in a remote cave in western Georgia, a few dozen miles east of the Black Sea shore, scientists on an archaeological dig were searching among scattered stalagmites for pieces of the past. Ancient bones were strewn about on the floor of the cave, but those held only mild interest for the team. Instead, they gathered buckets of sediment, on the hunt for ancient DNA.
Ancient DNA, like that extracted from the mummified remains of Copper Age humans and frozen mammoths, used to be a precious find — obtained only through the careful sequencing of nicely preserved fossils and bones. Finding the stuff was not easy and usually required a lengthy trek to the Arctic, a large research budget and a fair amount of luck. But now, scientists are finding it everywhere.
The results of the cave study, published this past April in Scientific Reports, showed that bears, roe deer and bats were present in this region at least as far back as 80,000 years ago. But finding traces of late Pleistocene animals is just scratching the surface of what can be done with environmental DNA, or eDNA, the traces of genetic molecules from long-dead organisms that survive as cell-free residues in the soil or other terrain. One of its strengths is that it can detect the remnants of organisms with soft bodies, allowing scientists to reconstruct entire ecosystems complete with plants, algae and more. Environmental genomics (or metagenomics, as it is also known) truly lets us see the ancient world in a few grains of sand.

Laura Parducci, an evolutionary plant ecologist at Uppsala University, uses eDNA to understand how plant communities have responded to climate change in past eras.
Courtesy of Laura Parducci
According to Laura Parducci, an evolutionary plant ecologist at Uppsala University in Sweden who was not involved with that paper, “The big benefit [of metagenomic techniques] is that you can get DNA from species that are actually not visible in the fossil records.” Parducci studies ancient ecosystems by extracting eDNA from sediments from lake beds in Scandinavia. She hopes to understand how plant communities responded to climate change in the past. Gleaning insights into ancient plants has traditionally lagged behind that of animals because plant remnants like seeds and wood typically decay more readily than durable animal bones and can be harder to find in the geologic record. Moreover, Parducci also wants to reconstruct the past of organisms such as microbes, which leave no obvious trace.
‘What Happens to That DNA?’
The moment an organism dies, its body begins to decompose. Its cells rupture, and their contents spill into the environment. What scientists have realized over the past two decades is that even though the physical structure of a body disappears, its DNA can last for centuries.
Eske Willerslev, an evolutionary geneticist at the University of Copenhagen, has found this DNA in the least likely places, including the soil underneath glaciers, in caves and more. His work has helped rewrite the natural histories of many locations around the globe by reconstructing ecosystems as much as 450,000 years old, and maybe older. He claims the idea came to him as a graduate student. “It was the autumn, and I saw leaves falling from the trees, and I saw a dog take a crap on the street,” he recalled, all of which set him to wondering: “‘What happens to that DNA? Could it somehow be preserved in the sediments or in the soil where it was put?’”

Eske Willerslev, an evolutionary geneticist at the University of Copenhagen, was one of the first to show the potential of remnant eDNA to reveal details of ancient ecosystems.
Mikal Schlosser
His graduate research, on microbes that had been trapped in ice sheets between 2,000 and 4,000 years ago, had required him to extract ancient DNA from glacial ice cores. But ice cores were expensive to obtain, and the idea that there might be other, more accessible places where ancient DNA had accumulated was appealing to him.
But his thesis adviser laughed when Willerslev brought up the idea because the prospects seemed so dim. Over time, DNA outside of cells accumulates various types of damage. For example, it gets broken down into tiny fragments, the ends of which get riddled with errors from nucleotide base-pair substitutions, further garbling whatever information it had encoded. The deep freeze of glacial ice seemed like the only place where DNA might stand a chance of surviving more or less intact for thousands of years.
Still, Willerslev pushed on. He laid hands on permafrost soil from an archaeological dig site, which would ordinarily have been discarded, and he managed to extract some DNA from it. The prevailing wisdom said that DNA should have been too riddled with base pair errors to say anything of value about the past.
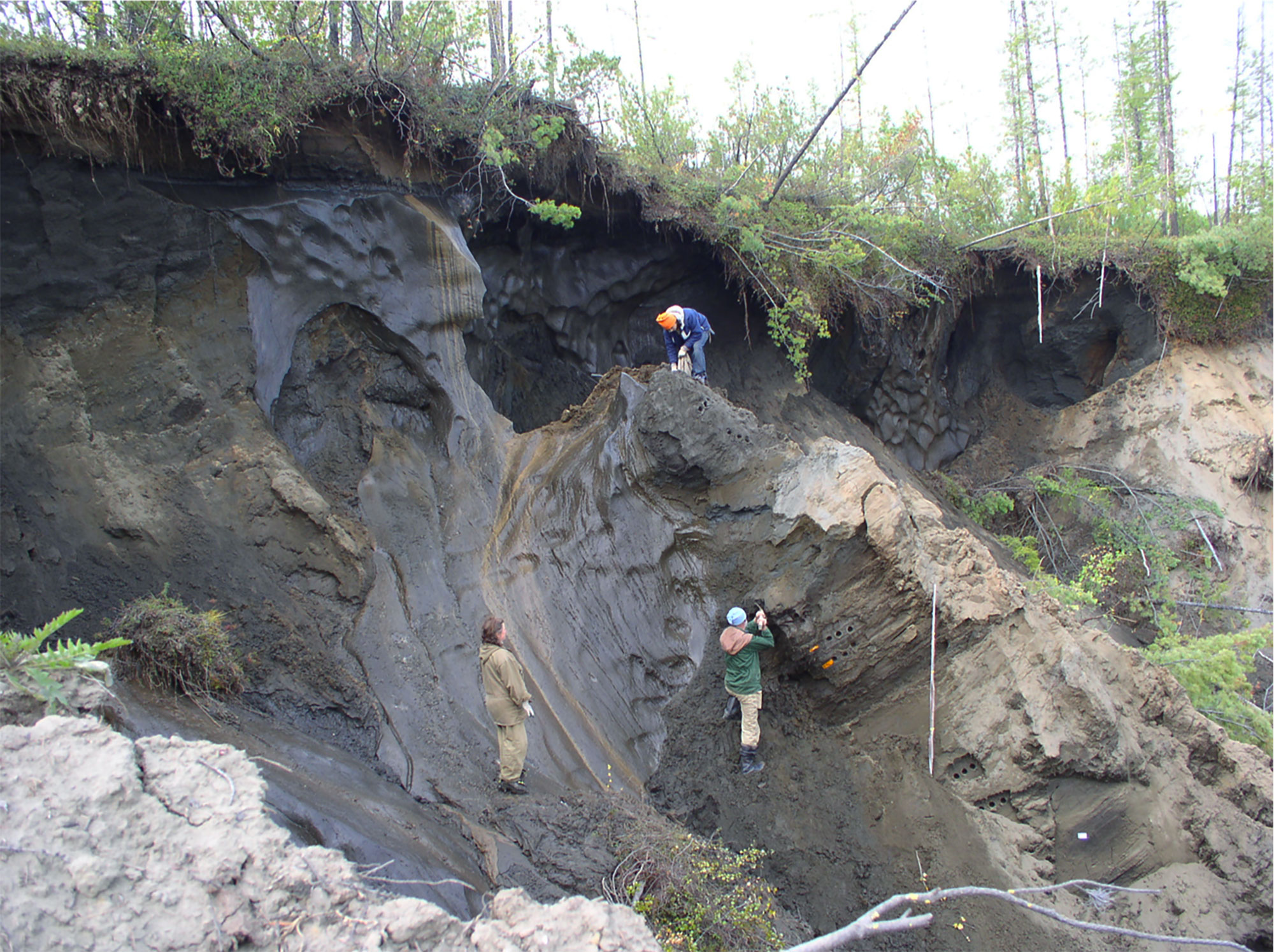

Willerslev’s team collects samples of permafrost sediments for eDNA extraction in northeastern Siberia.
Eske Willerslev
Instead, when he analyzed the sample, he was amazed to see that primers could pull out the DNA of ancient mammoths, bison, horses and a variety of plants. From just two grams of Siberian sediments dating back to between 10,000 and 50,000 years ago, he said, “you got this huge diversity.”
The study, published in 2003 in Science, was one of the first to showcase the potential of ancient environmental DNA. But the new method of obtaining the DNA necessitated a new analytical technique as well. “You can’t just take a DNA sample and sequence it as you would if you were working with a bone — it will be very messy,” Willerslev said.
Reconstructing the Past
A clump of ancient soil contains chopped-up bits of DNA from animals, plants and microorganisms that used to live in the area. Each species has a stretch of DNA letters unique to it. The challenge for scientists is parsing through the strings of As, Gs, Cs and Ts to identify their species of origin.
In a technique called metabarcoding, the DNA is extracted from the soil and either sequenced directly or amplified first to make more copies of it. Sometimes the scientists use universal primers, which recognize small sequences of DNA from all the organisms in one species or family; these molecules can then identify all the plant DNA or animal DNA to be amplified, or replicated, through the polymerase chain reaction. In another technique, called shotgun sequencing, all the DNA fragments are sequenced together simultaneously. Sequences isolated by either technique are then matched to a library of known sequences, and scientists can see what specific plants, animals or microorganisms are contained in the sample. Willerslev likens the process to police forensics, in which DNA obtained from a crime scene is collected and then matched to a database of criminals.
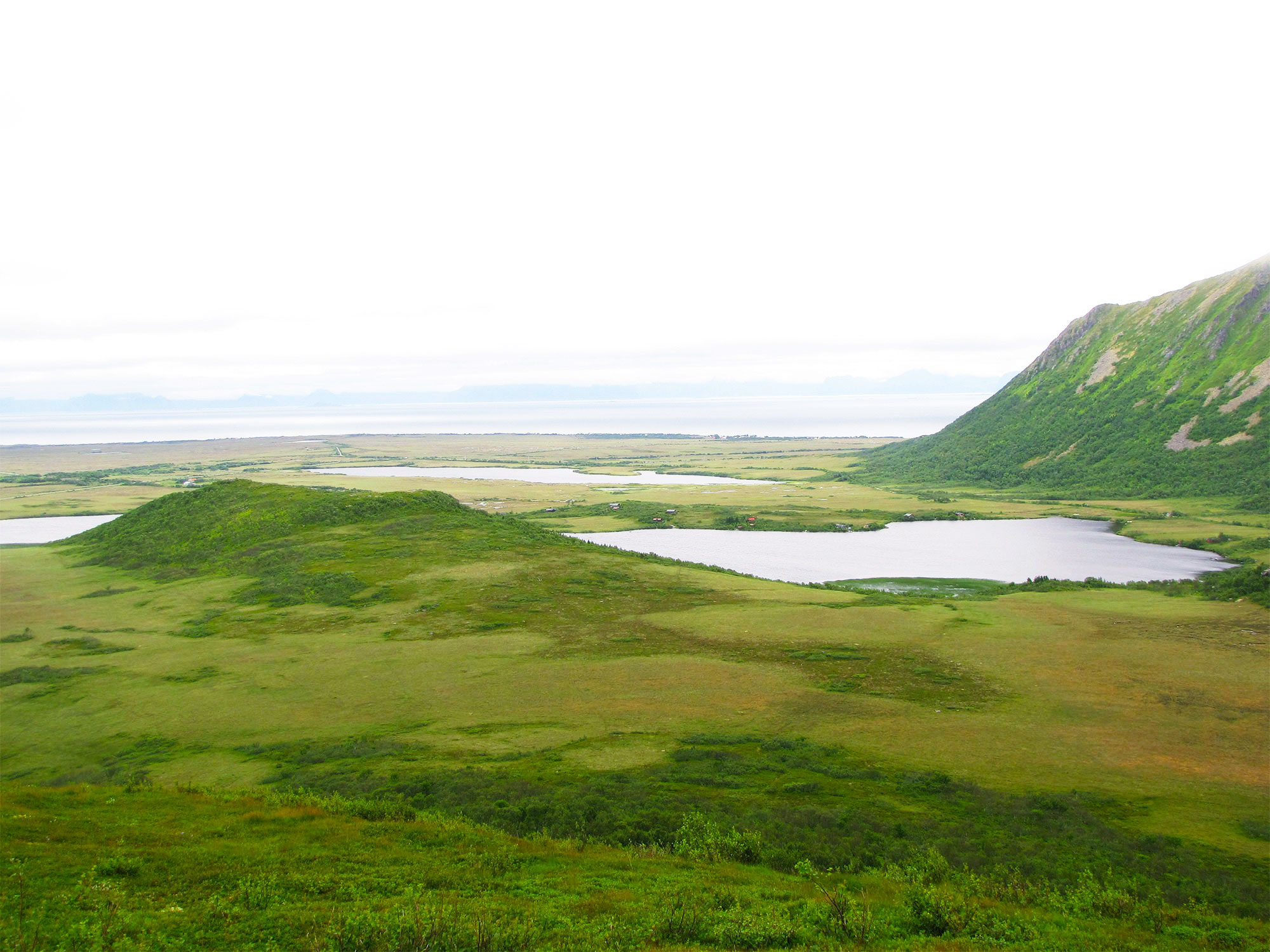

Sediments from lake beds like this one in Andøya, Norway, have yielded samples of DNA from diverse forms of life from thousands of years ago, most of which left no other surviving physical trace of their existence.
Courtesy of Laura Parducci
Creating an accurate picture of the past depends on the quality and preservation of the ancient DNA sample. When an organism dies, its DNA becomes vulnerable to a variety of threats, like DNA-digesting enzymes that spill out of the dead cells and microbes that feast on the decaying tissues. The environment can also be a threat, blasting the genetic material with UV rays. Over time, the DNA is fragmented into small pieces, and its sequence becomes riddled with nucleotide deletions and errors.
In a process called hydrolytic deamination, for example, the base cytosine spontaneously loses its amino group and becomes uracil. Normally in DNA, cytosine pairs with guanine, and thymine pairs with adenine. When the ancient DNA is replicated in the lab, this damage results in higher than normal adenine in the DNA strands. This process is so predictable in ancient DNA that scientists even look for it as a way to confirm the antiquity of the molecules.
Scientists prefer to find DNA in ice or permafrost because it slows down or inhibits these destructive processes. Caves and lake beds are protective environments, too. So far, the oldest eDNA recovered was from permafrost soil; dating techniques place it as somewhere between 450,000 and 800,000 years old. Samples any older than that are probably too hopelessly degraded and contaminated to be meaningful. So unfortunately for fans of Jurassic Park, dinosaur DNA — dating back tens or hundreds of millions of years — is never likely to be retrieved.
Parducci, who primarily studies ancient DNA trapped in lake beds, explains that there are many caveats when using eDNA. First, some of the primers used in metabarcoding work better than others. As a result, they they amplify their targeted fragments of DNA at different rates, which can distort the original DNA ratios.
Second, the databases for matching and identifying the extracted DNA are not complete. Databases, which are created when scientists upload DNA sequence data about individual species, do not reflect the total amount of biodiversity in the wild. Willerslev nonetheless expresses optimism that “the databases are growing exponentially, meaning the level of identification improves every year.”

Jamie Wood, a paleoecologist at Manaaki Whenua Landcare Research in New Zealand, notes that eDNA researchers must constantly be on watch against misleading contaminants. In some samples, he has found DNA from bananas, which clearly had to come from workers’ meals.
Bradley White/Manaaki Whenua
Jamie Wood, a paleoecologist at Manaaki Whenua Landcare Research who studies ecosystem function and composition in New Zealand over the past 50,000 years, explains that painstaking measures also have to be taken to ensure modern DNA doesn’t get mixed up in the ancient sample.
He says that it is easy to contaminate the ancient samples with modern DNA, and it can then be difficult to tease apart the two. He occasionally finds banana in his samples, for example, even though banana was not a plant native to New Zealand, where he collects eDNA. That banana was likely to have been introduced by scientists who ate the fruit and handled the samples, or by those who manufactured the reagents. Banana has also been sequenced a lot and is well represented in the plant DNA database, which increases the chance of spurious matches. In his work Wood has to be careful to weed out fact from fiction.
To do so, he combines his DNA analysis with information from pollen and geologic records. “Each one tells you something about the past,” Wood said. “When you use them all together, the real power comes in understanding those past ecosystems.”
DNA From an Ancient Pathogen
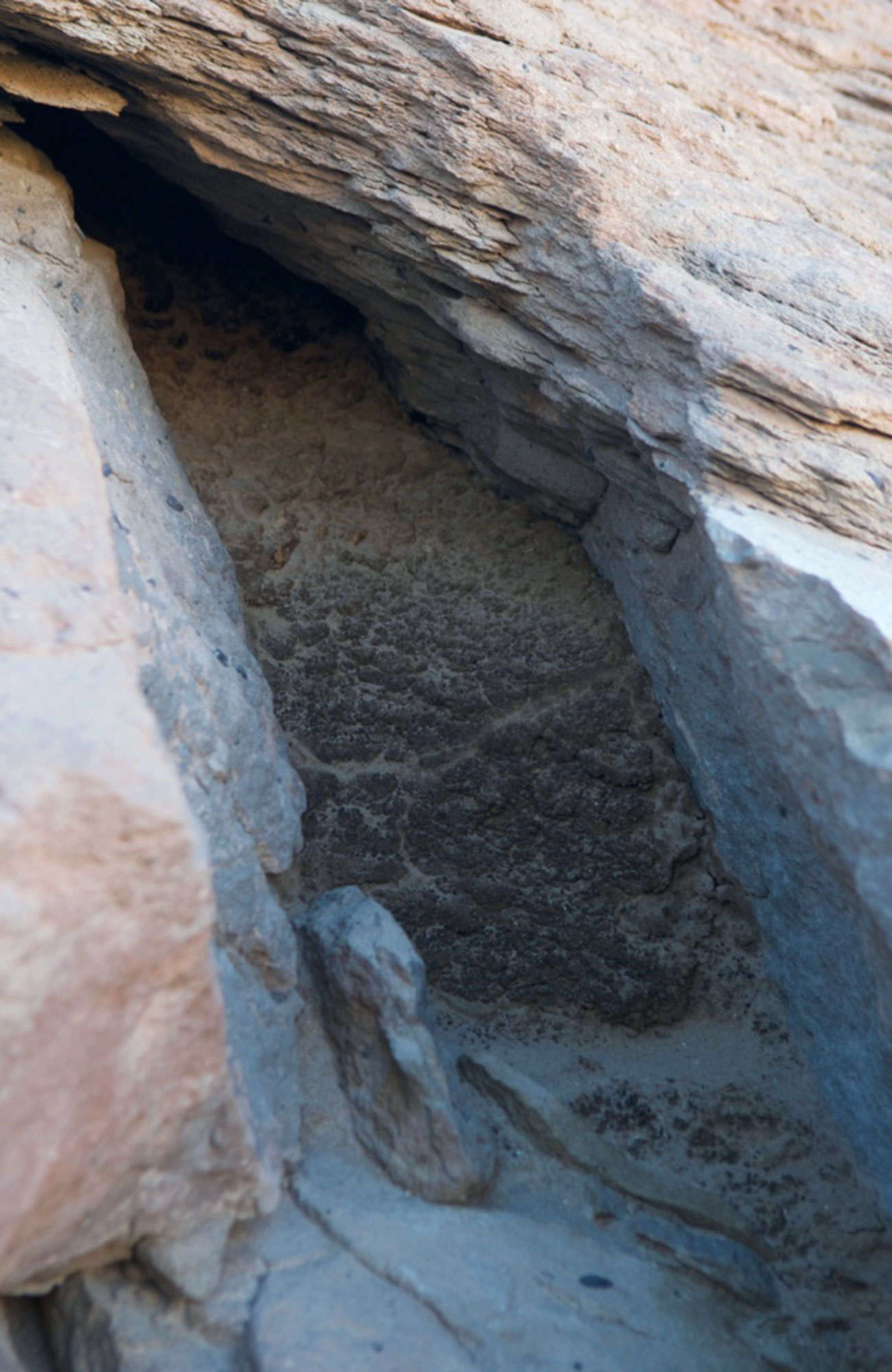

A rodent midden in the Atacama Desert sampled by Wood’s team. It may date back to 50,000 years ago.
Claudio Latorre
Wood recently obtained ancient DNA from rodent middens that were buried in rock crevices in the Atacama Desert in Chile from as far back as 50,000 years ago. The middens, made from bits of plants and covered in ancient rodent urine, held DNA that was well preserved. Wood was particularly interested in the plant pathogens he might find, and how their prevalence may have shifted after the last ice age, during a period when annual precipitation in the region was higher.
The results, published in Scientific Reports last November, found six different taxa that contained only plant pathogens. Although he could not identify the exact species present because the DNA was too deteriorated, he was able to narrow down his sample. In one example, he found oomycetes, a group of organisms called water molds, that include things like Phytophthora, the microorganism that causes potato blight.
The paper reports on what seems to be the oldest DNA recovered from plant pathogens. Wood found for some of these, increased rainfall drove their prevalence. He is interested in using data from the past to predict how plant diseases, especially those that affect crops, might be affected by a warming and wetter climate. For Wood, this type of work “could help improve models and [the] ability to predict how things might change with future climate change.”
Digging in the Reefs
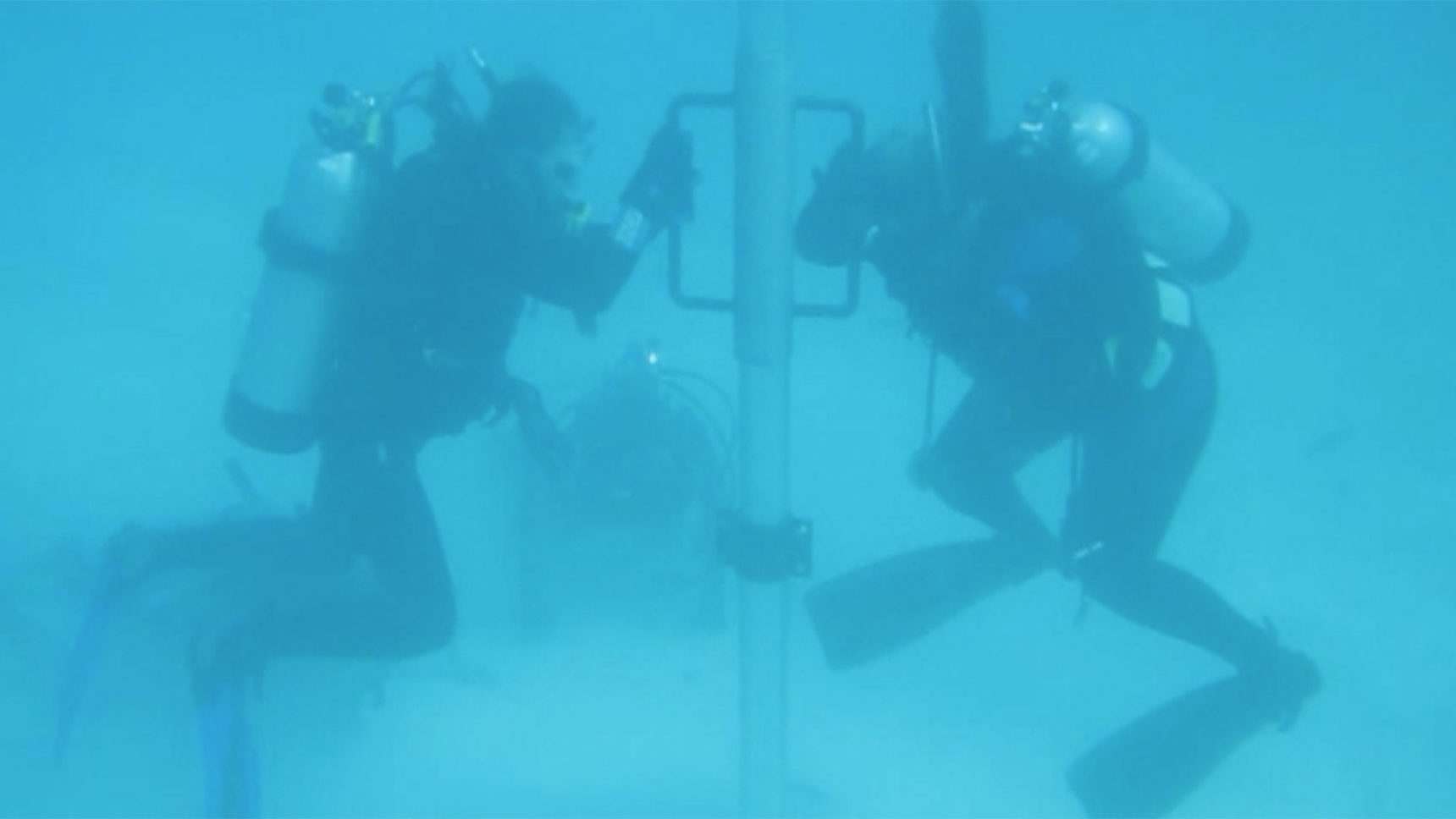
While many scientists are scouring caves, lakes and fossilized rodent droppings for ancient DNA, others are taking to the seas. The marine biologist María del Carmen Gómez Cabrera and her team of researchers from the University of Queensland in Australia are using ancient environmental DNA from the ocean floor to understand how the community of corals in the Great Barrier Reef changed over time.
Over the past 200 years or so, fast-growing corals in the Great Barrier Reef were replaced by a slower growing, pollution-tolerant variety. The change, which coincides with the rise of European colonization in Australia, became more pronounced during the past 70 years, a time marked by widespread water pollution from clearing the land, raising livestock and growing monoculture crops.
Coral reefs are wonderfully diverse ecosystems housing thousands of species, many of which do not leave a fossil record. For Gómez Cabrera, learning about the other organisms that share the Great Barrier Reef with corals would provide a wealth of information about the reef’s health over time. When corals die, from pollution in the water or the pathological condition called coral bleaching, for example, they leave an empty space that is quickly colonized by seaweed. Certain types of seaweed can accordingly be used as a proxy to gauge coral die-offs. Gómez Cabrera therefore set out to find signs of seaweed in the eDNA record of the Great Barrier Reef, as a guide to historical rates of coral death there.
“There was always this nagging problem in the back of our minds that we were trying to paint a picture of past ecosystems based only on very few players,” said Gómez Cabrera.
To obtain the sediment cores, she and her team donned scuba gear and headed into the Great Barrier Reef to collect samples of sediment. They hammered aluminum pipes, about five meters long and 10 centimeters in diameter, into the hard and cemented corals that had been deposited over the 13,000-year life span of the reef. “We just pound and pound and pound with the hammer into the sediment as far as we could get [the tube] in,” Gómez Cabrera said. “It’s very hard labor.”
After the cores were removed and taken back to the lab for analysis, the scientists obtained seaweed DNA from as far back as 750 years ago. In their paper, published in February, they found that the relative abundance of brown seaweed was great during periods when the corals were hard to find, and vice versa. The results indicate that brown seaweed DNA could be used to understand coral growth over time.
The authors caution that their results could be influenced by the differential preservation of ancient DNA, since the DNA from all organisms and locations might not be equally well protected against degradation over time. But the finding was replicated in many different cores throughout the reef, supporting their conclusions.
This early study demonstrates that eDNA can provide valuable information about the community structure of soft-bodied organisms in the coral reef. In the future, that might provide a more complete picture of reef health.
As the methods improve and the reference DNA libraries expand, many scientists are encouraged to think that environmental DNA will eventually allow them to glimpse not only past biodiversity but also past evolutionary change. Within a few years, “I think you can start looking at things like selection, for example,” Willerslev said. By observing gene frequencies over time in a species, they can glean a picture of how that organism evolved in response to environmental changes and selection pressures.
“Research on environmental DNA will greatly increase what we know about the past,” according to Mareike Cordula Stahlschmidt, an evolutionary anthropologist at the Max Planck Institute for Evolutionary Anthropology in Leipzig. Stahlschmidt, who was the first author of the stalagmite paper, thinks it is a powerful new addition to the paleoecology toolbox.
The scene is playing out at archaeological dig sites around the world: Scientists are increasingly interested in the piles of soil unearthed from archaeological digs, rather than just the bones and artifacts. If you imagine that one archaeological dig site can provide materials for dozens of labs at the fraction of the cost of obtaining bones, it is reasonable to say that the field will be expanding rapidly.
“I think this is where the future is,” Willerslev said.