Evolving Bacteria Can Evade Barriers to ‘Peak’ Fitness
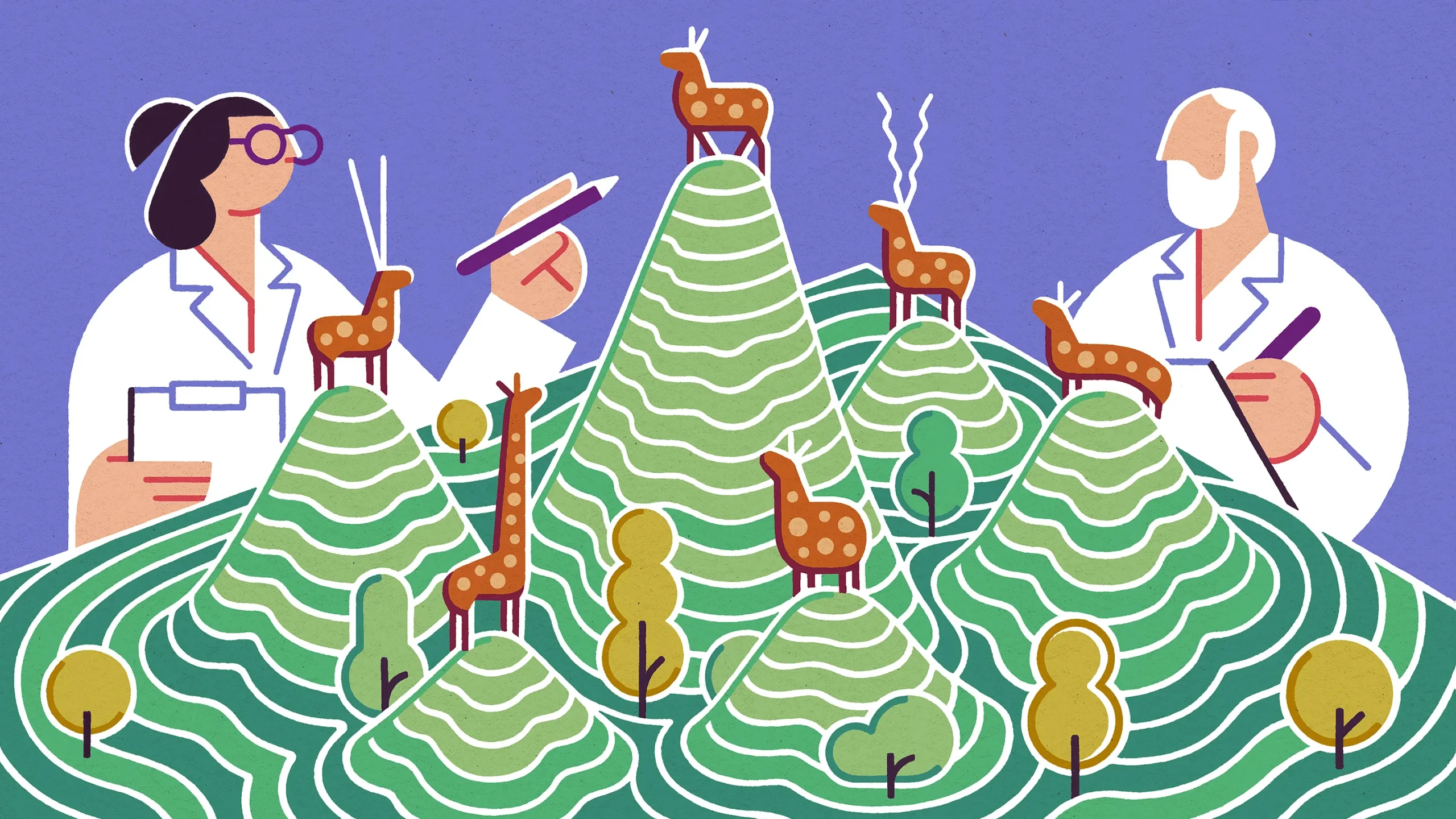

Researchers are learning new things about the “fitness landscapes” that they sometimes use to plot evolutionary trajectories for organisms.
Carlos Arrojo for Quanta Magazine
Introduction
Nearly a century ago, the evolutionary theorist Sewall Wright imagined a landscape of mountains and valleys. The peaks represented states of high evolutionary fitness for organisms, while the troughs between them represented states of low fitness. Organisms might move through the landscape by a process of mutation, climbing the peaks as their changing genes helped them achieve greater fitness.
Wright, a founder of modern population genetics, was intrigued by an apparent paradox: If a population of organisms managed to reach the top of a small hill, they would be marooned there, surrounded by worse states. They could not reach higher peaks without first traversing the doldrums below, something that natural selection would not normally allow.
Over the course of the last hundred years, evolutionary biologists have used mathematical models and, increasingly, lab experiments with living organisms to explore how populations of all sizes can move through fitness landscapes (sometimes called adaptive landscapes). Now, in a study just published in Science, researchers have engineered more than a quarter-million versions of a common bacterium and plotted each strain’s performance to create one of the largest lab-built adaptive landscapes ever. It enabled them to ask: How hard is it to get from any given point to the peaks?
Surprisingly, the rugged fitness landscape was traversable for most of the bacteria: About three-quarters of the strains had a feasible evolutionary route to antibiotic resistance. The findings support the idea, indicated by earlier theoretical work, that “valleys” in fitness may be more easily avoided than one might think. They also open the door to a better understanding of how real populations — of bacteria but also perhaps other organisms — might change under the pressure of natural selection.

The new paper from Andreas Wagner at the University of Zurich adds to his career of work on understanding fitness landscapes as tools of evolutionary biology.
Thi My Lien Nguyen for Quanta Magazine
For many decades, exploring fitness landscapes was primarily the reserve of theoreticians working with simulated organisms, or pioneering experimentalists working on a relatively small scale. But with the rise of easy, inexpensive gene editing technology, the team behind the new paper wondered if they could build a very large adaptive landscape using living organisms, said Andreas Wagner, a professor of biology of the University of Zurich and an author of the new paper.
They decided to plot the fitness effects of a single gene in the bacterium Escherichia coli. Dihydrofolate reductase, the enzyme that this gene encodes, is a target of the antibiotic trimethoprim, and mutations in the gene can make the bacterium resistant to the drug. Wagner and his colleagues, including lead author Andrei Papkou, a postdoc at the University of Zurich, created more than 260,000 genetically distinct strains of E. coli, each of which used a different permutation of nine nucleotides encoding three amino amino acids in the functional core of its version of the enzyme.
They grew the strains in the presence of trimethoprim and kept track of which ones thrived. The plot of their data revealed a landscape with hundreds of peaks of various heights, representing how well each of the genetic variants (genotypes) enabled the bacteria to evade the drug.
Then the researchers looked at how hard it would be for the different strains to evolve to reach one of the highest peaks. For each genotype, they calculated what series of mutations would be necessary to transform it into one of the highly resistant strains.
As Wright predicted decades ago, some paths ended atop low peaks that left no opportunity for further improvement. But many of the paths — routes by which, one mutation at a time, organisms could change their genotypes — reached fairly high points.
“We got good statistics on how frequently they get stuck on low peaks,” Wagner said. “And it’s not frequently at all. … Seventy-five percent of our populations reach clinically relevant antibiotic resistances.”
That tallies with what Sam Scarpino, a biologist and disease modeler who is the director of AI + Life Sciences at Northeastern University, said he would expect. “They have this very nice result that we’ve predicted,” he said, pointing to a recent theoretical paper exploring the relationship between the ruggedness and navigability of fitness landscapes. When fitness landscapes are high-dimensional — when they go beyond the simple three dimensions of most people’s imaginations to, say, the nine dimensions used in Wagner’s study — very different networks of regulatory genes that produce the same physical traits are more likely to be close together on a landscape or to be connected by an accessible pathway.
Merrill Sherman/Quanta Magazine
For example, Wagner and Papkou found that the highest peaks for antibiotic resistance in their experimental landscape were often surrounded by the nine-dimensional equivalent of very wide slopes; in effect, they were shaped more like Mount Fuji than the Matterhorn. As a result, many genotypes started out somewhere on the slopes of the highest fitness peaks, which made it easier for those strains to get to the top.
It wasn’t a given that the highest peaks would draw in the vast majority of genotypes, noted James O’Dwyer, a theoretical ecologist at the University of Illinois, Urbana-Champaign. But in this landscape that seems to have been the case.
That’s why building fitness landscapes as Wagner, Papkou and their colleagues did — massive ones based on real organisms — is an important step in bridging the gap between what we might assume to be true and what actually exists in nature, in systems far more complex than we can easily picture, said Ben Kerr, a professor of biology at the University of Washington. “How do we map our intuitions … onto situations that are not part of our experience?” he said. “One has to retrain one’s intuition. A good starting point is to do it on empirical data.”
Researchers at the University of Zurich created 260,000 genetically distinct strains of the bacterium Escherichia coli and evaluated their resistance to one antibiotic.
SERCOMI/Science Source
As huge as the fitness landscape in Wagner’s new paper is, it shows only what the bacteria are capable of in a single specific environment. If the researchers changed any of the particulars — if they changed the dose of the antibiotic or raised the temperature, say — they would get a different landscape. So although the findings seem to suggest that most E. coli strains can evolve antibiotic resistance, that outcome might be either far less likely or far more likely in the real world. All that seems certain is that most strains probably aren’t irrevocably sabotaged by their own minor successes.
Intriguing next steps for this research could therefore involve exploring whether any of the rules that seemed to prevail in the experiment’s version of the landscape might be more broadly universal. “If they were, there would be some underlying deep reason for it,” O’Dwyer said.
Wagner and Papkou are hoping to explore other versions of the landscape in future work. Papkou notes that it is not possible to map every permutation of even a single gene comprehensively — the landscape would explode to astronomical size almost immediately. But with lab-built landscapes and theoretical models, it should still be possible today to begin exploring whether universal principles undergird how an evolving entity can change in response to its environment.
“The bottom line is: It is pretty easy for Darwinian evolution to start in a suboptimal position and move by force of natural selection to a high fitness peak,” Papkou said. “It was pretty astonishing.”
Correction: December 4, 2024
An earlier version of the article incorrectly described how many amino acids and nucleotides were involved in the permutations generated for the new experiment.



