Insects Conquered a Watery Realm With Just Two New Genes
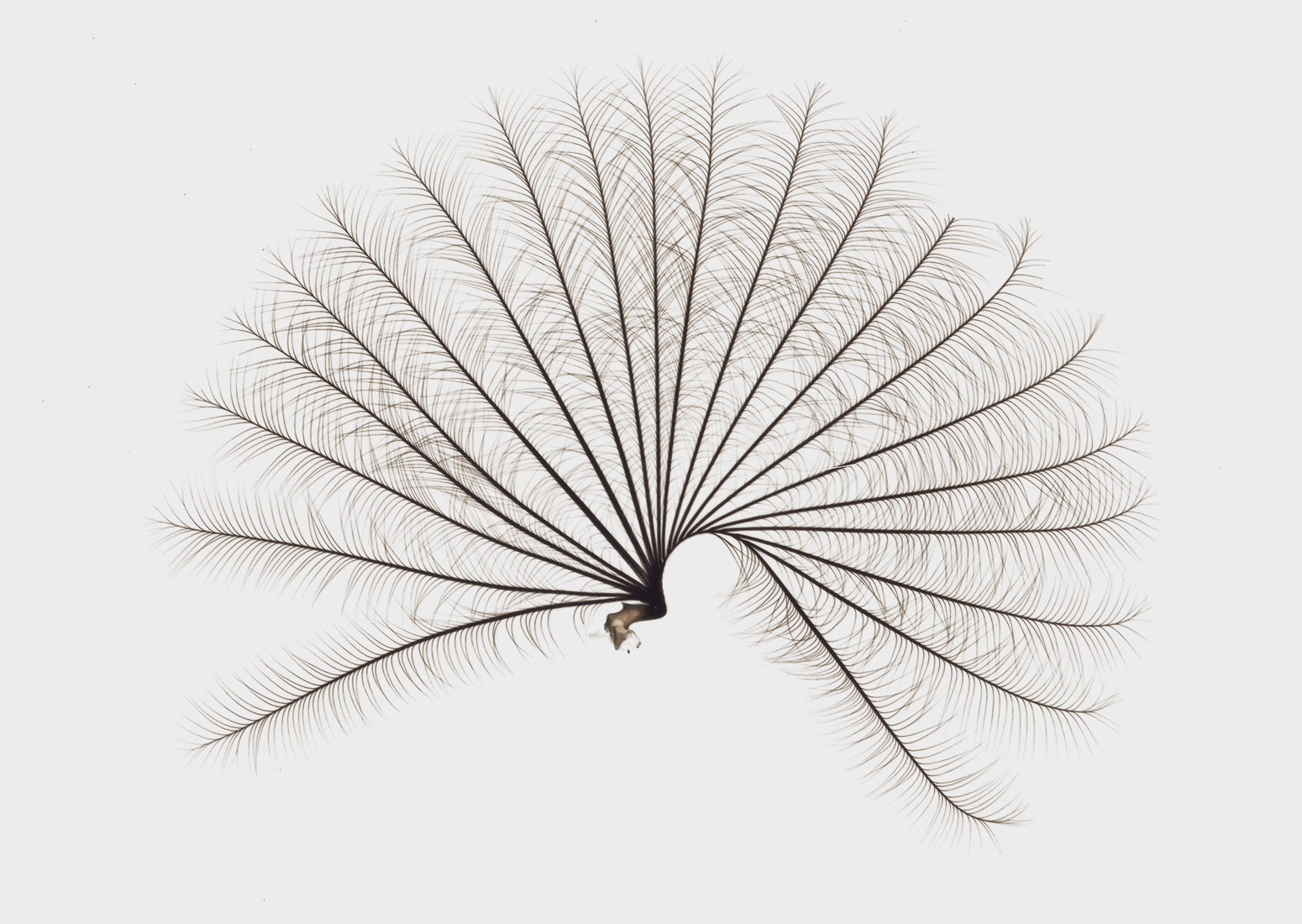

This delicate fanlike organ is a unique addition to the legs of one group of water striders.
Courtesy of Dr E. Santos and Dr A. Khila
Introduction
Ever since Darwin articulated his theory of natural selection, the question of evolutionary novelties has intrigued biologists. It’s relatively easy to understand how natural selection can reshape an existing trait — to make antlers bigger, legs longer or wings more colorful. But sometimes a fully formed trait appears seemingly out of the blue, without any apparent antecedent. Where did it come from?
Part of the answer can be found in a new study appearing today in Science that shows how the sudden emergence of just one or two new genes can profoundly transform organisms’ appearance, behavior and ecological niche. Using developmental genetics, evolutionary analysis, biomechanics and ecology, the researchers paint a picture of how a vital novelty evolved within one group of aquatic insects. But the significance of the discovery as a model for evolutionary innovation could extend throughout the animal kingdom.
“People have shown with comparative genomics that novel genes can be involved in novel structures. But this is the first time, to my knowledge, that the direct link is established from a novel gene to a novel structure to the invasion of a completely new ecological opportunity,” said Abderrahman Khila, an evolutionary and developmental genomicist at the Institute of Functional Genomics of Lyon, who led the study on the delicate insects called water striders.
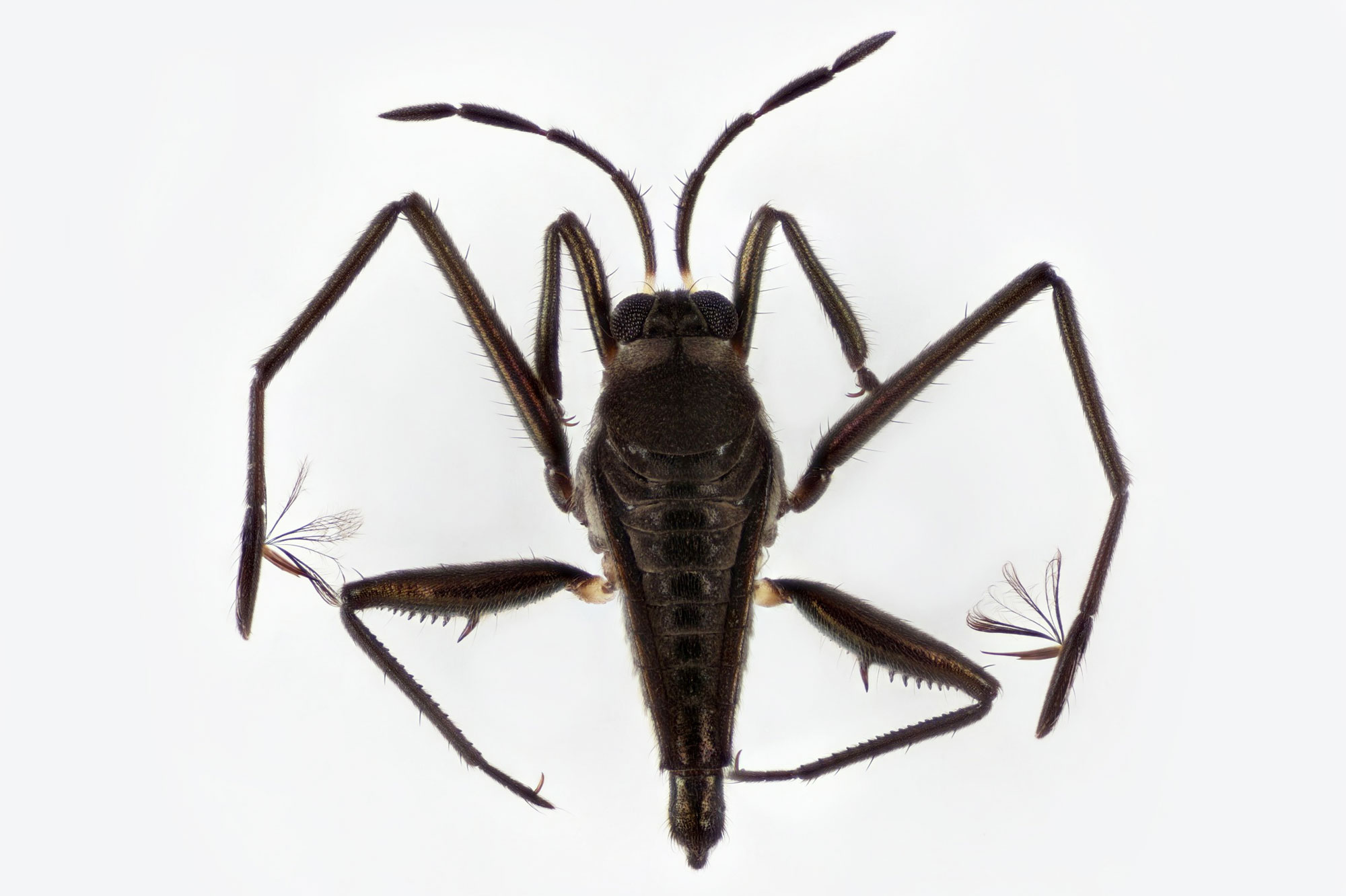
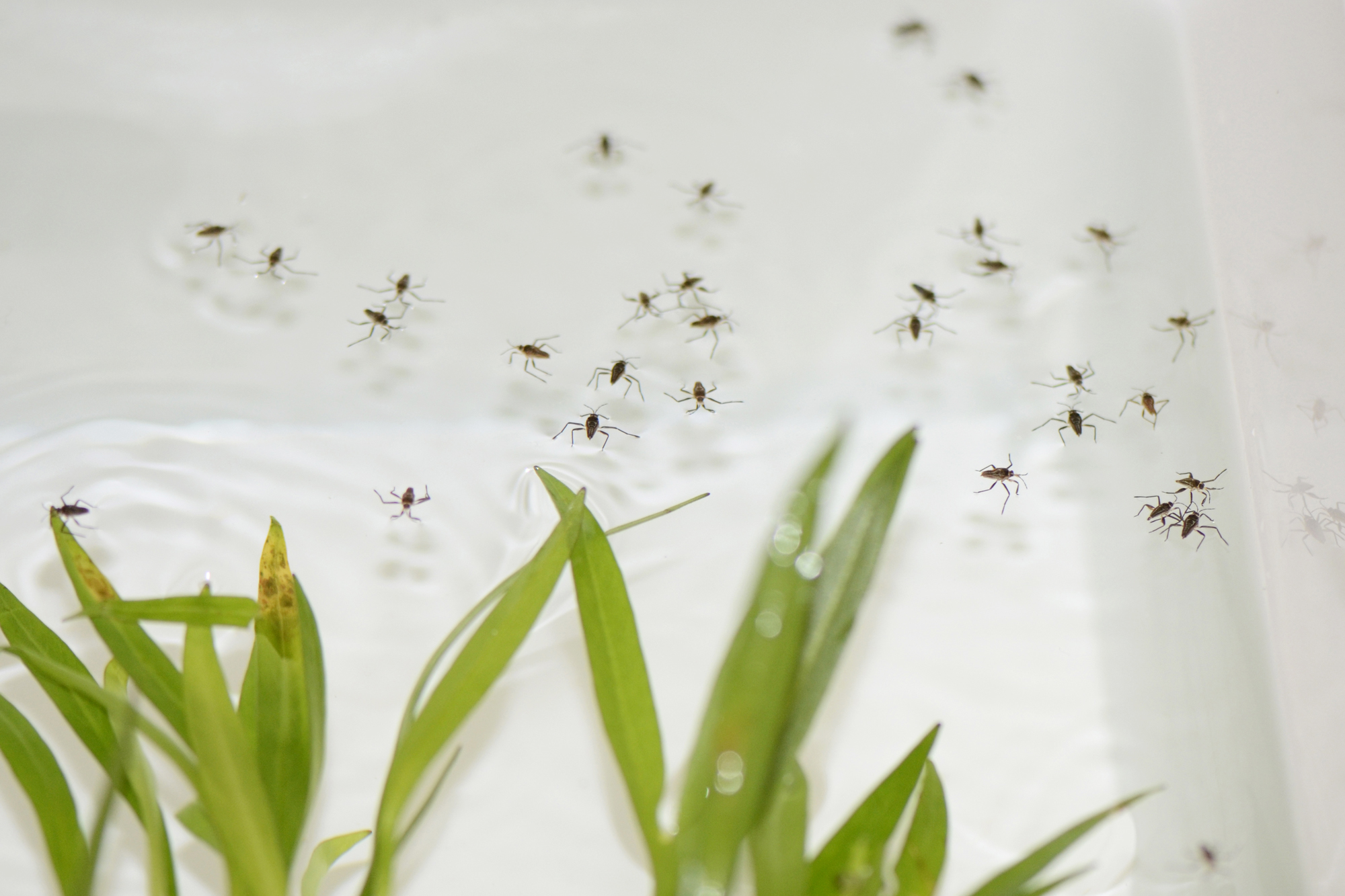


A water strider of the genus Rhagovelia (top) has a specialized structure on its middle pair of legs that water striders from other groups lack. This novel feature enhances the ability of Rhagovelia to navigate across fast-moving water.
Carnegie Museum of Natural History (top); Dr E. Santos and Dr A. Khila
A water strider of the genus Rhagovelia (left) has a specialized structure on its middle pair of legs that water striders from other groups lack. This novel feature enhances the ability of Rhagovelia to navigate across fast-moving water (right).
Carnegie Museum of Natural History (left); Dr E. Santos and Dr A. Khila (right)
Water striders are adept at navigating the surface of still water; they glide across ponds and lakes around the world. Those in the tropical genus Rhagovelia, however, have also figured out how to walk across fast-flowing streams and turbulent whitewater. Their secret asset is a special extendable structure on their middle leg resembling a Japanese fan, which no other water striders have. By deploying the fan, the insects can increase their leg’s contact with the water surface and can push against the water more forcefully. It’s a clever adaptation, but how did Rhagovelia acquire it when it has no precursors in other water striders?
To understand where the leg fan came from, Khila and his postdoctoral fellow Emília Santos, along with a couple of students, first had to figure out how to rear the water striders in the lab — a non-trivial task with what turned out to be a finicky species. It took about three years to figure out how to maintain the insects throughout the entire life cycle, from egg to adult. Once the colony was established, the team was ready for experiments.
Khila and Santos ground up the developing legs of the Rhagovelia water striders and sequenced the transcriptome — the complete suite of genes active in those tissues. The fan is only present on the second pair of legs, so the researchers compared the gene activity in the second pair to that of the first and third pairs. They discovered about 80-90 genes that were overexpressed only in the second legs.
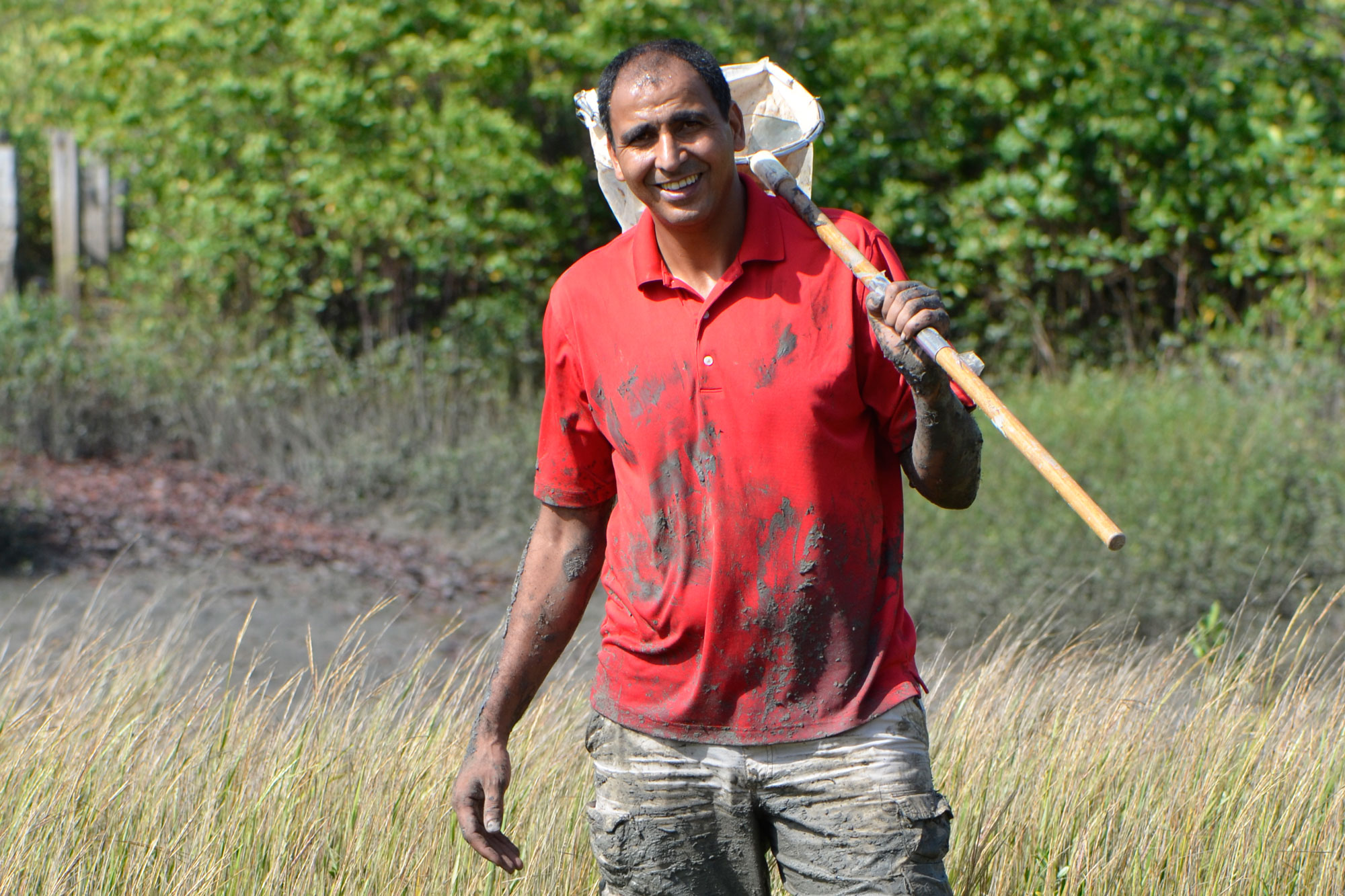

Abderrahman Khila, an evolutionary and developmental genomicist at the Institute of Functional Genomics of Lyon, studies how genomic changes in water striders correspond to their evolution of new capabilities. Here he is shown doing field work in Belem, Brazil.
Isabelle Cordeiro
Next, they used a method called in situ hybridization to pin down where in the leg those 80-90 genes were active. Khila and Santos identified five genes from that group that were expressed specifically in the tip of the second leg, where the fan develops. Three of the five were related to the structure of the cuticle, the protective outer layer of the insect’s exoskeleton. The other two genes appeared to be paralogs — genes that are the result of a duplication event in the DNA. The function of the paralogs was unknown.
In search of clues about the function of the two paralogs, the researchers looked for the genes across many water strider species. By looking at the evolutionary history of genes in the lineage of water striders, the researchers uncovered the ancestral copy of the gene (distinguished from the more recent duplicate) and nailed down the moment in evolutionary time when the duplicate appeared: at the origin of the Rhagovelia genus. Only water striders in the Rhagovelia genus have the duplicate gene, and they are also the only ones to have a leg fan.
“The evolution of the fan coincides with the duplication of that gene,” Khila said, also noting that the expression of the gene in the tip of the leg is significant. “It’s a big smoking gun.”
Because the leg fan reminded the researchers of a Japanese fan, they decided to call the newer gene geisha and its ancestral version mother of geisha. To learn what the genes were doing, they used a method called RNA interference to “knock down” (or turn off) the expression of those genes. Knockdown of geisha and mother of geisha caused the resulting insects to make only small, rudimentary fans. (Because of the strong sequence similarities between the two genes, the scientists have not yet been able to turn off one without the other, so differences between their functions are still unclear.)
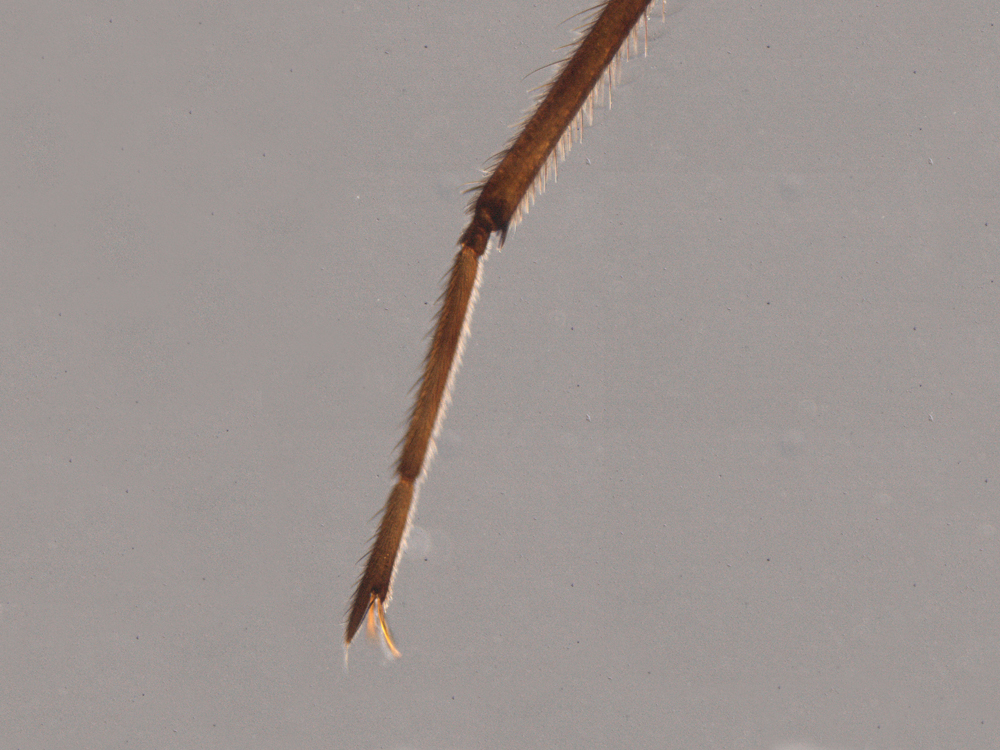
The second leg of a Stridulivelia water strider ends in a simple claw, while that of a Rhagovelia water strider also carries an extensible fanlike structure.
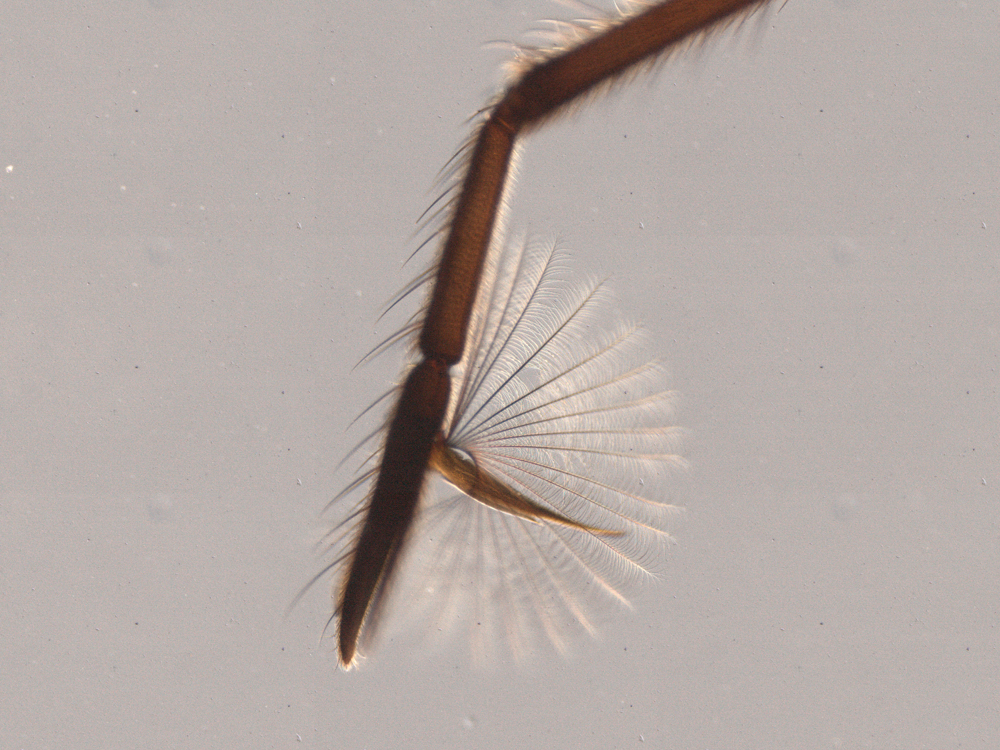

The fan improves the ability of Rhagovelia to push against the surface tension of water.
The second leg of a Stridulivelia water strider (left) ends in a simple claw, while that of a Rhagovelia water strider also carries an extensible fanlike structure. The fan improves the ability of Rhagovelia to push against the surface tension of water.
Dr E. Santos and Dr A. Khila
Khila and Santos then challenged the insects in the water to figure out what the fans were doing for them. They compared four groups: a closely related genus of water strider called Stridulivelia that does not have fans, untreated Rhagovelia, Rhagovelia in which the fan was surgically removed, and Rhagovelia in which geisha and mother of geisha were knocked down by RNA interference.
The researchers observed how the insects coped with two different watery environments. In calm water, the fan did not appear to provide a speed advantage in locomotion, although the water striders with fans could go farther with each stroke of the leg, which meant they could paddle more slowly. “So they can go just as fast without working as hard,” Khila said.
In the challenging environment of flowing water, however, the fans provided a crucial advantage: They enabled the insects to move upstream, while those without fans were swept away by the current. The Rhagovelia with knocked-down genes, which had developed only rudimentary fans, could cope with slowly moving water but failed at higher speeds. In short, the fans — even in a primitive form — were critical for enabling Rhagovelia species to navigate across running water, where closely related species could not.
Khila said it’s rare for experiments to be able to bring together biomechanics, transcriptomics, evolutionary biology, ecology and developmental genetics, but that combination is what’s needed to provide a complete answer to how phenotypic evolution occurs.
“The integrative, interdisciplinary nature of this demonstration is what’s striking here,” said Ehab Abouheif, an evolutionary developmental biologist at McGill University. (Abouheif was not involved in the study but was Khila’s postdoctoral adviser between 2006 and 2011.) According to Abouheif, Khila didn’t just show that this new gene makes the water striders’ fan; he also “reconstructs its evolutionary history, and then does the trademark ecological tests to show that it has an ecological function.”
“This study is really impressive,” agreed Greg Wray, an evolutionary developmental biologist at Duke University. “Usually something like this gets assembled over a period of time, maybe by different groups, and then there’s one last piece that links them together. It’s impressive to be able to go basically across the whole arc of the project in one go like this.
“The other thing I think is interesting is that there are not that many cases where people have been able to use gene expression, absent any genetics, to zoom in so precisely on a genetic change that causes an obvious trait,” he added. “So that’s pretty impressive as well.”
Emília Santos, a postdoctoral fellow in Khila’s laboratory, worked with him on identifying the genes responsible for Rhagovelia’s leg fan and superior control on moving water. Mélisandre Tefit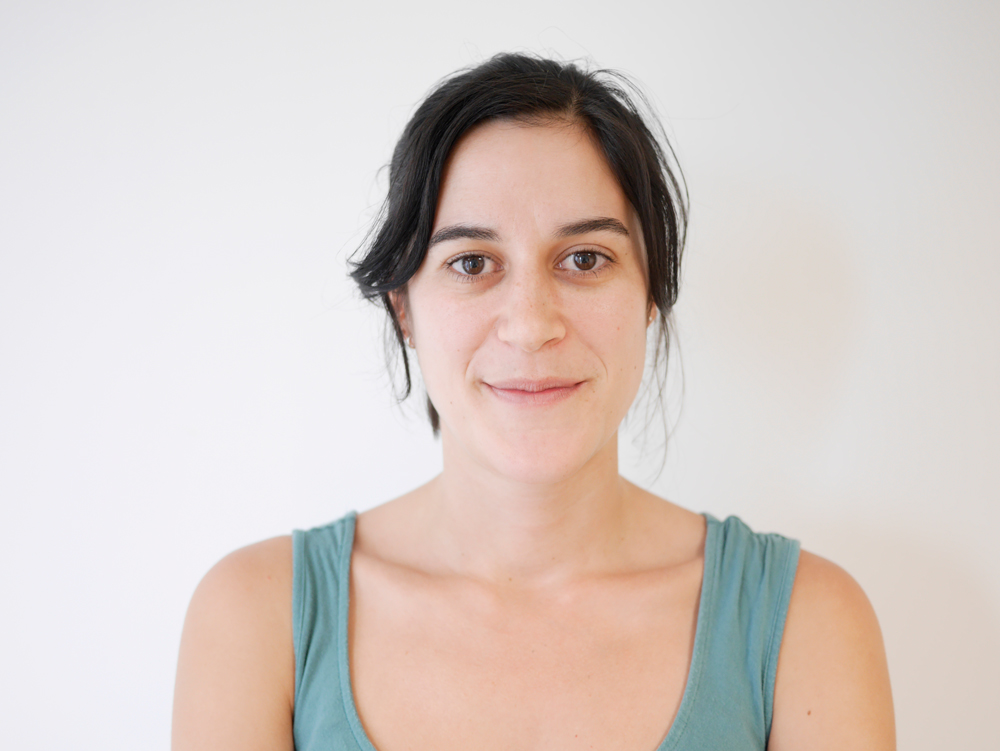
That said, biologists familiar with the study also find it interesting that geisha and mother of geisha are novel genes that don’t have a known role in development. “So far, the literature [about evolutionary novelties] is dominated by examples of co-option and repurposing of ‘old’ genes and pathways into ‘new’ contexts,” said Armin Moczek, an evolutionary developmental biologist at Indiana University, in an email.
“From butterfly-wing spots to beetles’ horns to numerous other examples, the image that emerges is that of organisms as Lego creations, as the modified reassemblies of the same and seemingly very limited pool of genes, developmental pathways and morphogenetic processes,” Moczek continued. “We now expect to see novelty made possible through co-option. What this paper shows is that new genes continue to matter.” Even between closely related species, he said, the sudden emergence of new genes can help to facilitate “ecologically significant innovation.”
Ian Dworkin, a geneticist who studies fly evolution at McMaster University, noted that in the literature of evo-devo research — which looks at the role of developmental mechanisms in evolutionary changes — examples of co-opted genes are abundant. What is unclear is whether that abundance reflects the frequency with which co-option occurs in nature or whether it is the result of ascertainment bias (systematic sampling errors introduced by how biologists study evo-devo). “It is nice to see a cool innovation that implicates a novel gene — it’s not simply a co-option story,” he said. “That’s why this is really exciting.”
Dworkin wonders whether inserting and activating geisha and mother of geisha in Stridulivelia, the fan-less sister genus of Rhagovelia, would be sufficient to trigger the development of the leg fan. Although the experiment would be very difficult to pull off in a non-model organism, “that would be the real kicker to demonstrate that that is the key gene for it,” he said.
Evolutionary scientists often get stuck in a rut by thinking of the co-option of old genes or the emergence of novel ones as exclusive alternatives, Abouheif said. But this study is a reminder that evolution can be a mix of both, through the recycling of the old and the addition of the new into networks that create novelty. The question for researchers to ponder, he said, is: “How do these networks get assembled?”
This article was reprinted on ScientificAmerican.com.



