Theory Suggests That All Genes Affect Every Complex Trait

Complex traits such as eye color are the products of many genes working in concert. Just how many genes may be an open question — but the answer could be all of them.
Ashley Mackenzie for Quanta Magazine
Introduction
The question most of genetics tries to answer is how genes connect to the traits we see. One person has red hair, another blonde hair; one dies at age 30 of Huntington’s disease, another lives to celebrate a 102nd birthday. Knowing what in the vast expanse of the genetic code is behind traits can fuel better treatments and information about future risks and illuminate how biology and evolution work. For some traits, the connection to certain genes is clear: Mutations of a single gene are behind sickle cell anemia, for instance, and mutations in another are behind cystic fibrosis.
But unfortunately for those who like things simple, these conditions are the exceptions. The roots of many traits, from how tall you are to your susceptibility to schizophrenia, are far more tangled. In fact, they may be so complex that almost the entire genome may be involved in some way, an idea formalized in a theory put forward last year.
Starting about 15 years ago, geneticists began to collect DNA from thousands of people who shared traits, to look for clues to each trait’s cause in commonalities between their genomes, a kind of analysis called a genome-wide association study (GWAS). What they found, first, was that you need an enormous number of people to get statistically significant results — one recent GWAS seeking correlations between genetics and insomnia, for instance, included more than a million people. Second, in study after study, even the most significant genetic connections turned out to have surprisingly small effects. The conclusion, sometimes called the polygenic hypothesis, was that multiple loci, or positions in the genome, were likely to be involved in every trait, with each contributing just a small part. (A single large gene can contain several loci, each representing a distinct part of the DNA where mutations make a detectable difference.)
How many loci that “multiple” description might mean was not defined precisely. One very early genetic mapping study in 1999 suggested that “a large number of loci (perhaps > than 15)” might contribute to autism risk, recalled Jonathan Pritchard, now a geneticist at Stanford University. “That’s a lot!” he remembered thinking when the paper came out.
Over the years, however, what scientists might consider “a lot” in this context has quietly inflated. Last June, Pritchard and his Stanford colleagues Evan Boyle and Yang Li (now at the University of Chicago) published a paper about this in Cell that immediately sparked controversy, although it also had many people nodding in cautious agreement. The authors described what they called the “omnigenic” model of complex traits. Drawing on GWAS analyses of three diseases, they concluded that in the cell types that are relevant to a disease, it appears that not 15, not 100, but essentially all genes contribute to the condition. The authors suggested that for some traits, “multiple” loci could mean more than 100,000.
The reaction was swift. “It caused a lot of discussion,” said Barbara Franke, a geneticist at Radboud University in the Netherlands who studies attention deficit hyperactivity disorder (ADHD). “Everywhere you went the omnigenic paper would be discussed.” The Journal of Psychiatry and Brain Science did a special issue just of response papers, some of them taking exception to the name, some saying that after all it was just an expansion of earlier ideas. A year on, however, the study has been cited more than 200 times, by papers whose subjects range from GWAS data to individual receptors. It seems to have encapsulated something many people in the genomics community had been turning over in their minds. But exactly what scientists should do with its insights depends on whom you talk to.
An Infinity of Small Effects
The origin of the idea lies in a very simple observation: When you look at the portions of the genome that GWAS findings have flagged as significant to individual traits, they are eerily well-distributed. Pritchard and his colleagues had been studying loci that contribute to height in humans. “What we realized was that the signal for height was coming from almost the whole genome,” he said. If the genome were a long string of ornamental lights, and every DNA snippet linked to height were illuminated, more than 100,000 lights would be shining all the way down the string. That result contrasted starkly with the general expectation that GWAS findings would be clustered around the most important genes for a trait.
Then, while looking at GWAS analyses of schizophrenia, rheumatoid arthritis and Crohn’s disease, the researchers found something else unexpected. In our current understanding, disease often arises because of malfunctions in key biological pathways. Depending on the disease, this might lead to the overactivation of immune cells, for example, or the underproduction of a hormone. You might expect that the genetic loci incriminated by GWAS would be in genes in that key pathway. And you’d expect those genes would be ones used specifically in the types of cells associated with that disease: immune cells for autoimmune diseases, brain cells for psychiatric disorders, or pancreatic cells for diabetes, for instance.
But when the researchers looked at disease-specific cell types, an enormous number of the regions flagged by GWAS were not in those genes. They were in genes expressed in nearly every cell in the body — genes doing basic maintenance tasks that all cells need. Pritchard and his colleagues suggest that this manifests a truth that is perhaps not always taken literally: Everything in a cell is connected. If incremental disruptions in basic processes can add up to greatly derange a trait, then perhaps nearly every gene expressed in a cell, no matter how seemingly unrelated to the metabolic process of interest, matters.
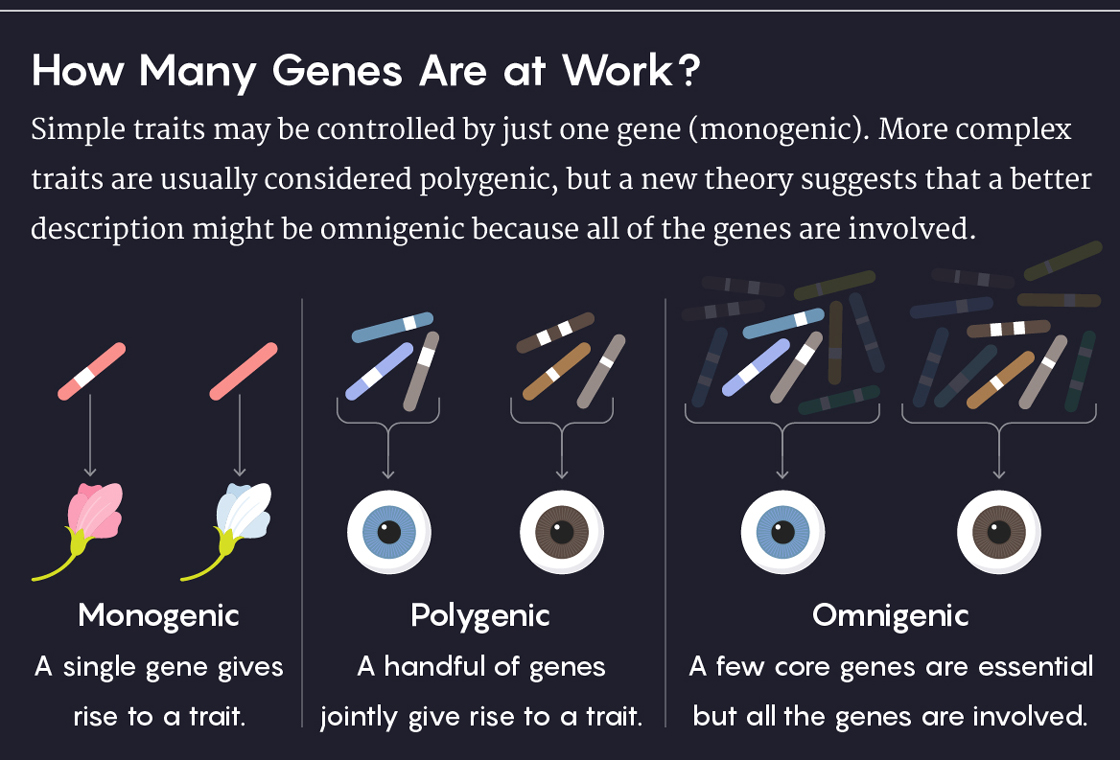

Lucy Reading-Ikkanda/Quanta Magazine
In its broadest strokes, this idea has been around since 1918, when R. A. Fisher, one of the founders of population genetics, proposed that complex traits could be produced by an infinite number of genes, each with infinitely small effects. But his was a statistical model that didn’t refer to any actual, specific biological conditions. It seems we are now in the era of being able to provide those specifics.
“This was the right paper at the right time,” according to Aravinda Chakravarti, a professor of neuroscience and physiology and director of the Center for Human Genetics and Genomics at New York University, who was a prepublication reviewer of the omnigenics paper in Cell. He and others had noticed many examples of how widely distributed genetic influences could be, he said, but they had not put them together into a coherent thesis. He disagrees with critics who say the paper simply stated the obvious. “The paper clarified many points of view. It didn’t matter if I had thought about it — I had not thought about it hard enough. And I had never heard anybody thinking about it hard enough, with any clarity, [such] that it formed any new hypothesis.”
In the paper, Pritchard and his colleagues proposed that, when geneticists seek what’s responsible for a disease or trait, it may be fruitful to think of the genes in a cell as a network. There may be some very highly connected genes at the center of a disease process, which they dub core genes. Peripheral genes, meanwhile, in aggregate help tip the scales one way or the other. The Cell paper authors suggest that understanding of the core genes will offer the best insights into the mechanism of a disease. Piecing together how peripheral genes contribute, on the other hand, will broaden understanding of why some people develop a disorder and others don’t.
Do Core Genes Exist?
Since the Cell paper’s publication a year ago, scientists’ discussion has circled around whether such a distinction is useful. David Goldstein, a geneticist at Columbia University, is not sure that disease processes must truly be routed through core genes, but he also says that the idea that not everything picked up by GWAS is central and specific to a given disease is important. In the early days of GWAS, he said, when a connection between a genetic locus and a disease was detected, people would take that as a sign that it should be the target of investigation for new treatments, even if the connection was weak.
“Those arguments are all fine — and were — unless something like what Jonathan is describing is going on,” he continued. “That’s a really big deal in terms of our interpretation of GWAS,” because weakly connected loci might then be less useful for getting at the pathology of a disease than people thought.
Yet that may well depend on the disease, according to Naomi Wray, a quantitative geneticist at the University of Queensland who pointed out when scientists first started doing GWAS analyses that they should expect to see many weak associations. A few conditions, she says, are primarily attributable to a small number of identifiable genes, or even just one — yet other genes may still flip the switch between one manifestation of illness and another. She cites the example of Huntington’s disease, a progressive neurological disorder caused by a specific defect in one gene. The age at which it strikes depends on how many repeats of a particular DNA sequence someone has in that gene. But even among patients with the same number of repeats, the age at which symptoms first appear varies, as does the severity with which the disability progresses. Scientists in the field are looking at other loci linked to Huntington’s disease to see how they might be causing the differences.
“These [loci] are by definition in peripheral genes. But they’re actually how the body is responding to this major insult of the core gene,” Wray said.
For most complex conditions and diseases, however, she thinks that the idea of a tiny coterie of identifiable core genes is a red herring because the effects might truly stem from disturbances at innumerable loci — and from the environment — working in concert. In a new paper out in Cell this week, Wray and her colleagues argue that the core gene idea amounts to an unwarranted assumption, and that researchers should simply let the experimental data about particular traits or conditions lead their thinking. (In their paper proposing omnigenics, Pritchard and his co-authors also asked whether the distinction between core and peripheral genes was useful and acknowledged that some diseases might not have them.)
Teasing out the detailed genetics of diseases will therefore continue to require studies on very large numbers of people. Unfortunately, in the past year, Pritchard has been told that some groups applying for funding to do GWAS have been turned down by reviewers citing the omnigenics paper. He feels this reflects a misinterpretation: Omnigenics “explains why GWAS is hard,” he said. “It doesn’t mean we shouldn’t do GWAS.”
Franke, who sees the paper as a provocatively phrased extension of earlier ideas, says that it has nevertheless shaped her thinking in the past year. “It made me rethink what I know about signal transduction — about how messages are relayed in cells — and how functions are fulfilled,” she said. The deeper you look at the workings of a cell, the more you realize that a single common protein may have quite different effects depending on what type of cell it is in: It may bear different messages, or block different processes, so much so that traits that might seem to be quite disconnected begin to change.
“It gave a lot of food for thought,” she said of the paper, “and I think that was the goal.”
This article was reprinted in Spanish at Investigacionyciencia.es.