Networks Untangle Malaria’s Deadly Shuffle
“Think of a deck of cards,” said Dan Larremore. Now, take a pair of scissors and chop the 52 cards into chunks. Throw them in the air. Card confetti rains down, so the pieces are nowhere near where they started. Now tape them into 52 new cards, each one a mosaic of the original cards. After 48 hours, repeat.
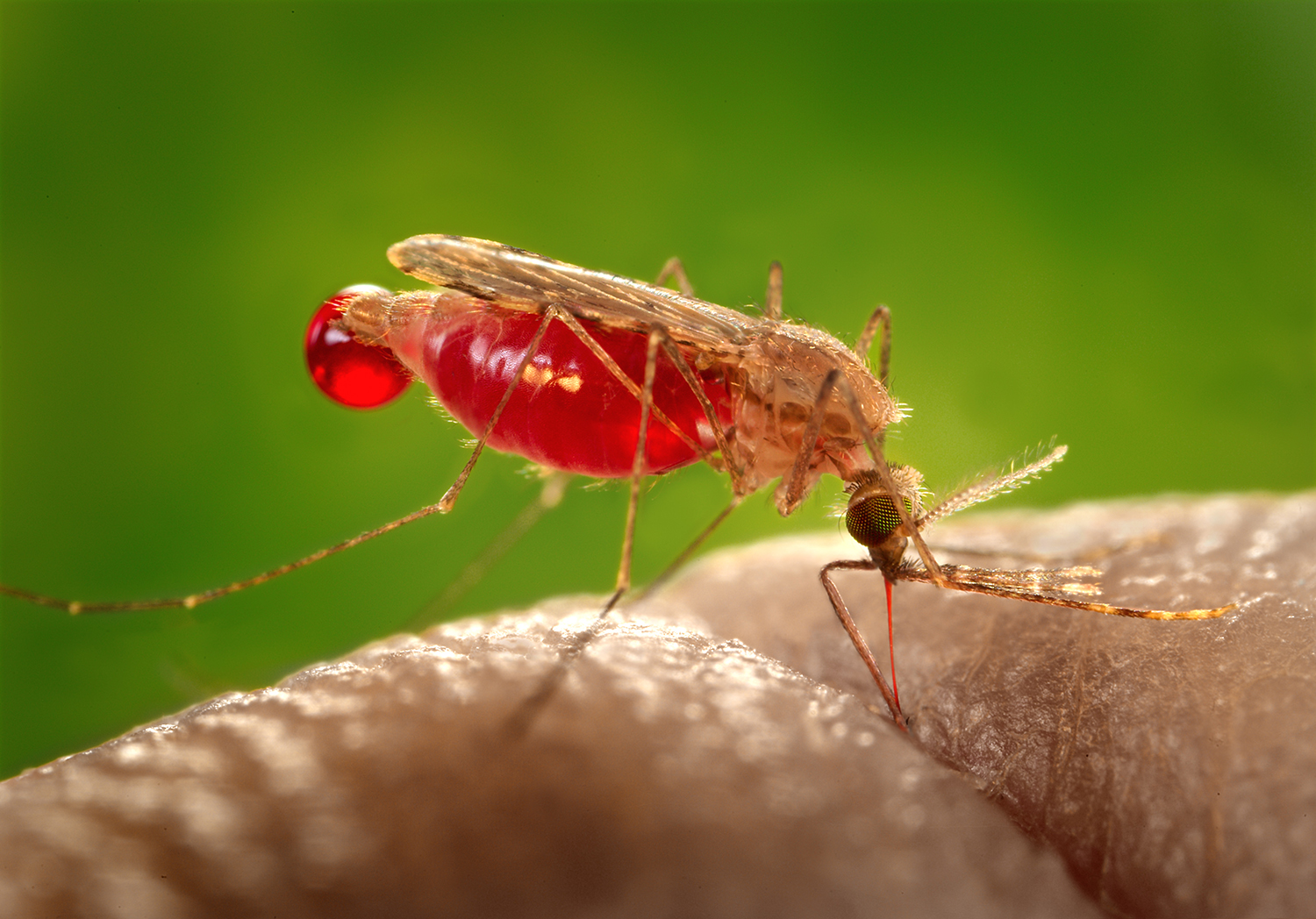
You have just reenacted the process that Plasmodium falciparum uses to avoid the immune system. P. falciparum is the world’s most dangerous malaria parasite, causing 600,000 deaths every year and killing more children under the age of 5 than any other infectious disease on the planet. Larremore, an applied mathematician, was introduced to its promiscuous habits while doing postdoctoral research at what is now the Harvard T.H. Chan School of Public Health.
Each card represents a gene for a protein that attaches to the walls of the host’s blood vessels, anchoring the parasite so that it cannot be dragged into the spleen, where it would be detected and destroyed. Each falciparum parasite has 50 to 60 of these var genes, as they are called, and as time passes the parasite uses first one, then another, presenting a constantly morphing face to immune cells that might spot it clinging to the blood vessel. The crowning glory of this tactic, though, is that when the parasite divides, which it does every couple of days, chunks and snippets of the genes swap places up and down the chromosomes. In one out of every 500 parasites, this process will generate an entirely new gene. With the number of parasites out there, that adds up quickly. “It’s crazy. It means the total number of var gene sequences in the world is millions and millions — virtually infinite,” said Antoine Claessens, a malaria researcher with the Medical Research Council, The Gambia Unit, in Fajara.
New evidence from Larremore and his collaborators, however, reveals a paradoxical stability in these genes. In a recent paper in Nature Communications, they show that while var genes themselves are never repeated, short sequences of DNA in them — pieces of cut-up card — are shared between species that have been separate for millions of years. It’s a finding that has made some malaria researchers feel hopeful, because it suggests that there are limits on the crazy remaking of the var genes, which could mean that vaccines can be developed to fight them.
Slicing and Dicing
“We want to know basic stuff,” said Caroline Buckee, an epidemiologist at the Harvard T.H. Chan School of Public Health and a co-author of the new study. “Are there certain parasites that cause disease more than others? Are they evolutionarily related to each other? … These questions, which in most pathogens we can figure out how to answer, have no answer [in malaria], because we don’t know how to compare these genes to each other.”
The usual tool for such a task is a phylogenetic tree. At the tree’s base is the oldest version of a gene, and as its daughters accrue small differences — a single DNA base change here, a single base change there — they become separate branches. Trees are built by lining the genes up next to each other and checking for differences at each DNA base. The trees have been helpful in studying the divergence of genes in viruses like the flu, which changes via just such a process of mutation.
Malaria researchers have used them as well, but with mixed results. A pair of var genes might have a chunk of 30 DNA bases in common, but if that chunk is at the beginning of one gene and at the end of another — which happens all the time in the shuffling process — a tree will call it a difference instead of a commonality. If the chunk does happen to be in the same place in both genes, a tree will say that the genes recently diverged, but the chunk could just as well have arrived two days ago in one gene and a year ago in another. All of this means that trees built from var genes are at best difficult to interpret, and at worst misleading, implying relationships where none exist. “It’s a mush. That’s the technical term,” joked Martine Zilversmit, a malaria researcher at the American Museum of Natural History in New York City.
If you want to compare these genes, though, there aren’t many other options. “It was a case of ‘this is the tool we have,’ and everyone kind of jamming their data into the tool,” said Buckee, who first began to talk about an alternative approach with collaborator Aaron Clauset, now a professor of computer science at the University of Colorado, Boulder, when the two were postdocs.
In 2012, Larremore, now at the Santa Fe Institute in New Mexico, took a postdoc position with Buckee and Clauset to try and see whether network analysis, a field he knew well, could help provide an alternative way to track the history of malaria parasites. Network analysis involves drawing links between nodes that have something in common, generating a diagram that can reveal underlying patterns. Linked nodes might be people who are friends in a social network, diseases that afflict the same people, or genes that share chunks of sequence.
If you make a network in which var genes are only connected if they share chunks of a certain length, the commonalities leap out. Buckee, Larremore and Clauset published a paper in 2013 showing that such networks could pick out identical sequences shared by P. falciparum parasites from different continents. Being able to clearly see these relationships helps researchers in their efforts to figure out how and why they arose. A greater number of common chunks in a pair of genes could mean that they share a recent ancestor, or it could indicate that the proteins they produce have a similar way of interacting with the immune system. The chunks could also be evidence of an ancestral cache of cut-up card pieces that modern parasites still carry around.
To investigate whether var genes exist in other parasite species and, if they do, whether they share any chunks with P. falciparum, the researchers analyzed samples of malaria parasites from wild apes. They used parasites from feces collected in the jungle and from the blood of sanctuary chimps, assembling DNA sequences from five Plasmodium species that infect gorillas and chimps, including one that had already been found to have var genes.
They were in luck: The var gene marker they searched for cropped up in at least three of the species. That in itself is interesting, because it means the var gene family is old — ancient, even, according to Thomas Lavstsen, a biologist at the University of Copenhagen in Denmark, who studies the genes but was not involved in the research.
When the team created their networks, they saw something else striking. The var genes shared chunks of their most variable regions with each other, despite the millions of years of evolution — millions of years of slicing and dicing — that separated them. The chimp parasite P. reichenowi in particular had so many connections to P. falciparum that in many places in the var genes, they are indistinguishable. “In other words,” Claessens said, “if I give you a [var gene marker], you cannot tell me if it comes from reichenowi or falciparum.”
This is important because it means that for millions of years, the var genes have been preserving the genetic variety produced by their snip-a-thon, rather than just chopping themselves up indiscriminately. There’s a good reason for them to do that, it turns out: As soon as the immune system starts to recognize one bit of a var gene, the parasite trots out newer versions. But after a while, the old bit will be rare enough that the immune system no longer recognizes it, and if it has been stashed away, rather than destroyed, the parasite can bring it back out again. Basically, this discovery shows that these old chunks “are not allowed to sink into evolutionary obscurity,” Zilversmit said.
The preserved pieces are also a sign that the genes’ diversity has limits, Lavstsen suggested. If the var protein changes too much, it can no longer cling to blood vessels. The fact that these chunks have stuck around all these years shows that they are part of the select range of structural options within which the protein still works. The new research sketches the borders of an evolutionary space beyond which var genes cannot venture without losing their function. “This has implications about how scared we should be about sequence diversity” as researchers work on developing vaccines, Lavstsen said.
As any malaria vaccine researcher will tell you, finding a method to that particular madness would be welcome. “There have been hardcore malaria vaccine projects going on for 40, 45 years,” Zilversmit said. “And it’s been really slow going.” Vaccines expose the immune system to a fragment of a pathogen protein that remains the same from infection to infection, so that the immune system will attack when it next encounters the protein. With most viruses, it’s a reasonable approach. With malaria, it’s a nearly impossible one. Even the shared sequences revealed in the recent paper are not repeated frequently enough to be useful targets for vaccines, Lavstsen said. “We’re trying to unscrew this Phillips-head screw with a flat-head screwdriver. It’s just not the right tool,” Zilversmit said.
Somehow, many people in malaria-endemic countries acquire natural immunity to the disease by adolescence. Researchers even know that their immune systems do it by recognizing the var-gene proteins. The difficulty is in figuring out how to do it artificially.
Lavstsen thinks that the overall shape of the protein, which is likely less changeable than the genes that produce it, may be what the immune system is recognizing in conferring natural immunity. This could be a better vaccine target than any particular sequence, he said. To adapt the deck-of-cards analogy, priming the immune system to recognize the rectangular shape of a card, rather than the patterns on its face, may have potential.
Peter Bull, a malaria researcher at the University of Cambridge, adds that one of the things he finds most remarkable about the research is the perspective it gives on the history of the var family of genes in the great apes. “This is a nice illustration of how this gene family has been around in the primate population for a very, very long time,” he said. “It fits in with the idea of all these new humanlike species that have been identified in the fossil record. There may have been a lot more primates very similar to us at a certain point. Perhaps they all had this kind of parasite.”
The Efficient Killer
The growing realization that var genes are not limited to P. falciparum has prompted researchers to reevaluate the genes’ gruesome powers. The dogma has long been that var genes are the harbingers of serious illness. Other human parasites that cause malaria, like P. vivax, do not have the genes and also provoke less severe disease. “Falciparum was unique,” Zilversmit said. “This was the mythology we built around it.”
But when it became clear some years ago that a chimp parasite had var genes, researchers began to question whether they had the right idea. And the new research reinforces the idea that the story may be more complicated than just the presence or absence of these genes. One of falciparum‘s worst tendencies is to latch onto the small blood vessels of the brain with var gene proteins. It sends sufferers, mostly children and pregnant women, into comas from which they do not wake. From the limited amount researchers know about malaria in other great apes, it isn’t clear that this cerebral malaria occurs in apes infected with var-gene-bearing parasites, even though the genes appear to produce the same kind of protein.
“It changes the kind of question you need to ask,” Zilversmit said. “Is there something about falciparum‘s var genes in particular that makes them bad? Maybe it has to do more with the humans themselves — maybe it has everything to do with the host and not with the parasite. In truth, it’s probably a little bit of both of those things. … [But] we’re going to have to look more closely to find out.”
Larremore and Buckee hope to use their knowledge of networks to answer other questions about the genes. How might exposure to malaria parasites when a person is young affect immune response to those var genes, for instance? By building networks in which the nodes could be anything from patients to var genes, it may be possible for researchers to observe revealing patterns. That work is still embryonic — “We are still working on assuring ourselves that we should believe the signal that we would get out,” Larremore said — but he is looking forward to seeing where it takes them. “I do think this is another opportunity for networks to deliver.”
This article was reprinted on ScientificAmerican.com.