Secrets of Early Animal Evolution Revealed by Chromosome ‘Tectonics’

Blocks of linked genes can maintain their integrity and be tracked through evolution, new research has shown. The discovery is the foundation of what is being called genome tectonics.
Runbo Chen for Quanta Magazine
Introduction
Chromosomes, the bundles of DNA that star in the mitotic ballet of cell division, play a leading role in complex life. But the question of how chromosomes came to exist and evolve has long been discouragingly hard to answer. This is due partly to the lack of chromosome-level genomic information and partly to the suspicion that eons of evolutionary change have washed away any clues about that ancient history.
Now, in a paper appearing today in Science Advances, an international team of researchers led by Daniel Rokhsar, a professor of biological sciences at the University of California, Berkeley, has tracked changes in chromosomes that occurred as much as 800 million years ago. They identified 29 big blocks of genes that remained recognizable as they passed into three of the earliest subdivisions of multicellular animal life. Using those blocks as markers, the scientists deduced how the chromosomes fused and recombined as those early groups of animals became distinct.
The researchers call this approach “genome tectonics.” In the same way that geologists use their understanding of plate tectonics to make sense of the appearance and movement of continents, these biologists are reconstructing how various genomic duplications, fusions and translocations created the chromosomes we see today.
The work heralds a new era in comparative genomics: Previously, researchers studied collections of genes from different lineages and described the changes one base pair at a time. Now, as more chromosome-level assemblies become available, researchers can trace the evolution of entire chromosomes back to their origin. They can then use that information to make statistical predictions and rigorously test hypotheses about how groups of organisms are related.
Two years ago, using similar innovative methods, Rokhsar and his colleagues resolved a long-standing mystery about the timing of genome duplications that accompanied the emergence of jawed vertebrates. But the importance of the approach isn’t purely retrospective. In the process of making these discoveries, the researchers are learning about the algebraically simple rules that govern what happens when chromosomes swap chunks of themselves. That information can guide future genomic studies and help biologists predict what they will find in the genomes of species that have not yet been sequenced.
“We’re starting to be able to get a broader picture of chromosome evolution across the tree of life,” said Paulyn Cartwright, a professor of ecology and evolutionary biology at the University of Kansas. Now, she said, scientists can draw conclusions about what was in the chromosomes in the very first animals. They can also examine how the various contents of the chromosomes changed or stayed the same — and why — as animals diversified. “We really could not do that before we had these high-quality genomes.”
What Ancient Genomes Share
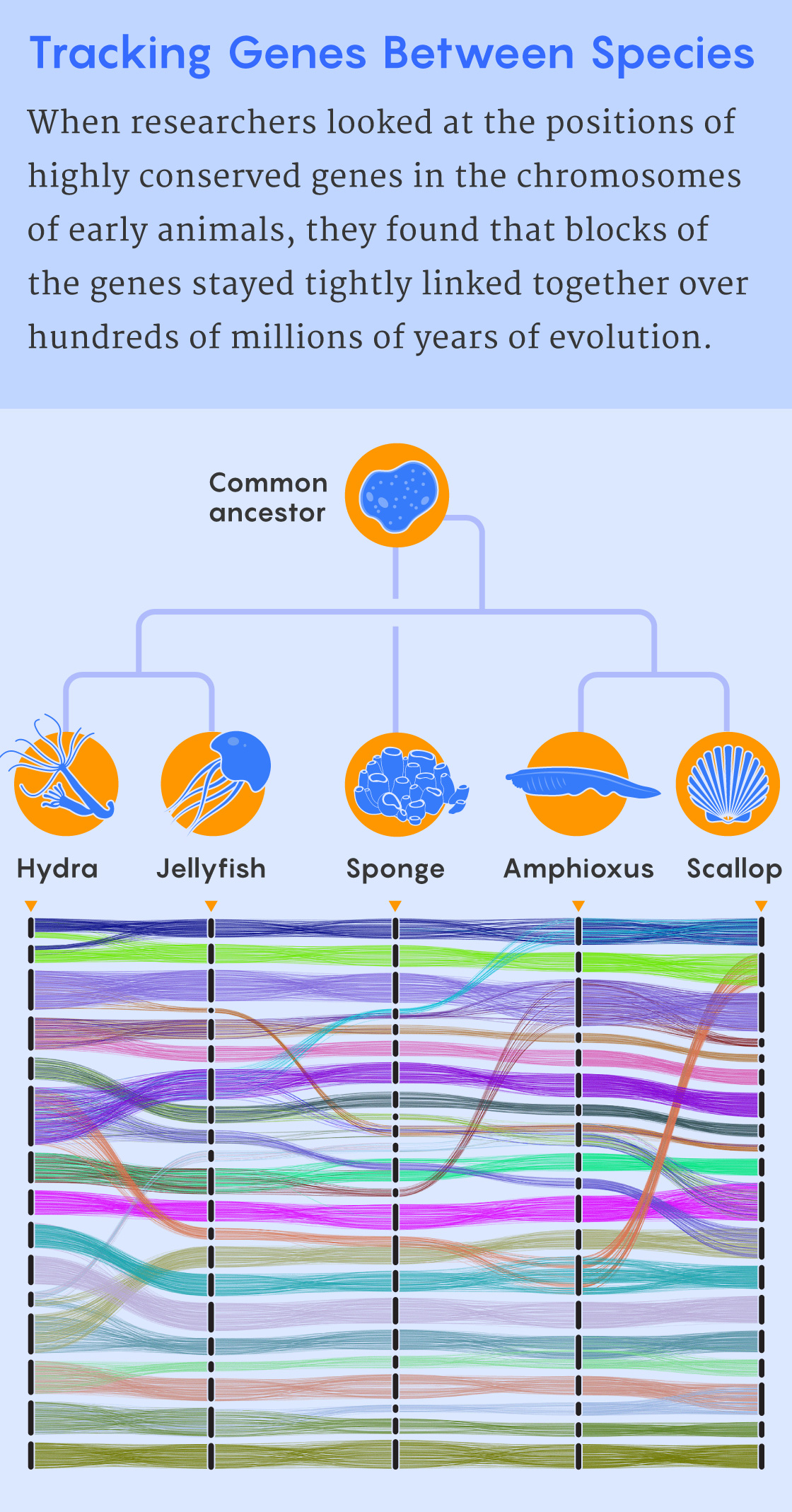
For the study published today, Rokhsar and a large international team of collaborators produced the first high-quality chromosome-level assembly of the genome of the hydra, which they described as a model for a “venerable cnidarian.” By comparing it to other available animal genomes, they discovered highly conserved groups of linked genes. Although the order of the genes within a block was often shuffled, the blocks themselves were stable over long stretches of evolutionary time.

Daniel Rokhsar, a professor of biological sciences at the University of California, Berkeley, suspected that blocks of genes might stay closely linked over millions of years of evolution, despite reshufflings in the order of those genes.
Mark Joseph / UC Berkeley
When scientists started sequencing animal genomes about 20 years ago, it was not a foregone conclusion to many of them that linked groups of genes on chromosomes could stay stable and recognizable over eons, let alone that it might be possible to track the passage of those blocks of genes through essentially all animal lineages.
Animals diverged from their unicellular relatives 600 million or 700 million years ago, and “being able to recognize the pieces of chromosomes that are still conserved after that period of time is amazing,” said Jordi Paps, an evolutionary biologist at the University of Bristol in the United Kingdom.
“Before we had this whole-chromosome data, we were looking at small snippets of the chromosomes, and we were seeing lots of rearrangements,” Cartwright said. “So we were assuming that there wasn’t conservation, because the genes themselves within a region of the chromosome are changing positions quite frequently.”
Still, although the order of genes was frequently reshuffled along chromosomes, Rokhsar intuited from his earlier studies of animal genomes that there was relative stability in which genes appeared together. “If you compared sea anemone or sponge to human … the fact that the genes were on the same piece of DNA seemed to be conserved,” Rokhsar explained. “And the pattern suggested that whole chromosomes were also conserved.” But that notion couldn’t be tested until recently, when enough chromosome-scale genomic information about diverse animal groups became available.
Genomic Inertia
But what would cause blocks of genes to stay linked together? One explanation for this phenomenon, which is called synteny, relates to gene function, says Harris Lewin, a professor of evolution and ecology at the University of California, Davis who studies the evolution of mammalian genomes. It may be more efficient for genes that work together to also be physically located together; that way, when a cell needs to transcribe genes, it doesn’t have to coordinate transcription from multiple locations on different chromosomes.
This probably does explain the conservation of some sets of genes whose arrangement is critical: The Hox genes that establish animals’ body plans, for example, must be in a specific order to set up body patterning correctly. But those tightly linked genes fall within a relatively short bit of DNA. Rokhsar says that he doesn’t know of any functional correlation that extends over a whole chromosome that could account for their findings.
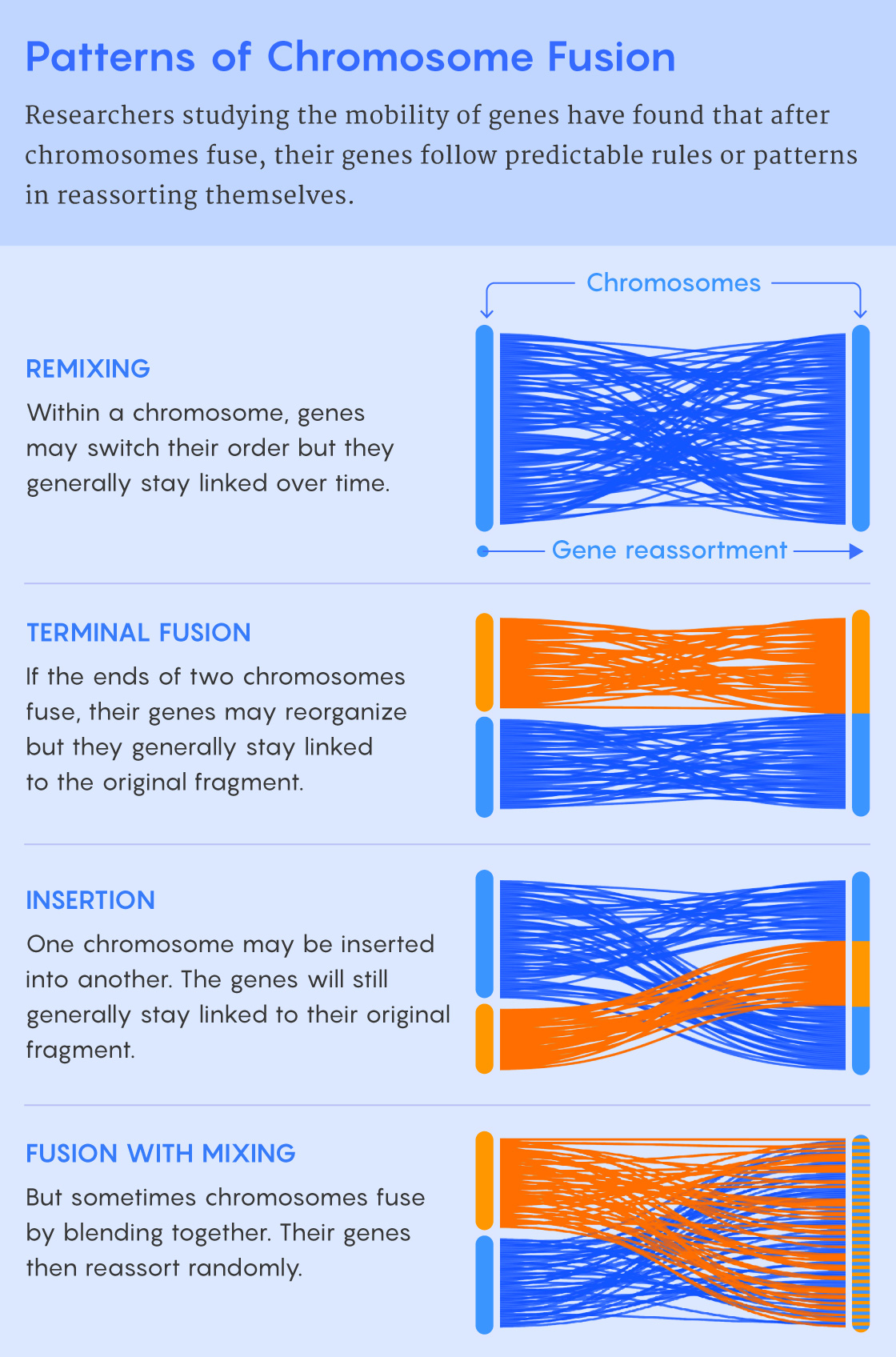
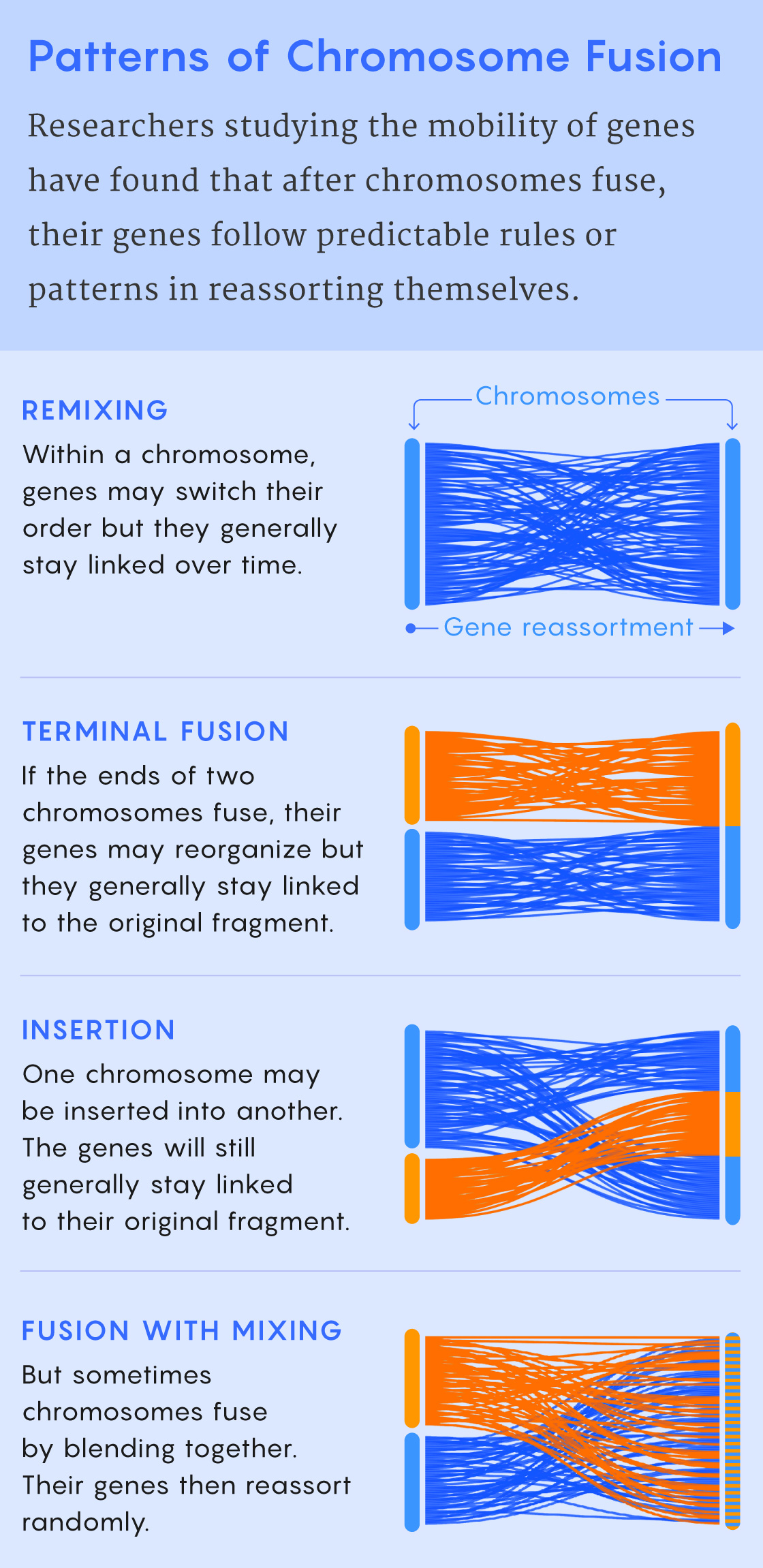
Samuel Velasco/Quanta Magazine; source: 10.1126/sciadv.abi5884
That’s why Rokhsar is skeptical of the functional explanation. It’s attractive (“That would be the coolest result, in a way,” he said) but also maybe unnecessary because unless a chromosome rearrangement conveys a big functional advantage, it’s inherently hard for the rearrangement to spread. And rearrangements are typically not advantageous: During meiosis and the formation of gametes, all chromosomes need to pair up with a matching partner. Without a partner, an odd-sized chromosome won’t become part of a viable gamete, so it is unlikely to make it into the next generation. Small mutations that reshuffle the gene order within chromosomes can still occur (“There’s probably a little room for error in terms of minor rearrangements, so they can still recognize each other,” Cartwright said). But broken or fused chromosomes tend to be dead ends.
Perhaps in groups like mammals, which have small population sizes, a rearrangement could randomly spread through what’s known as genetic drift, Rokhsar suggests. But in big, freely mixing populations like those of the marine invertebrates that spawn hundreds or thousands of eggs, “it’s really hard for one of the new rearrangements to take hold,” he said. “It’s not that they’re not being tried. It’s just that they never can get a toehold in evolution.”
Consequently, genes tend to stay stuck on one chromosome. “The processes by which they move are just slow, on a scale of 500 million years,” Rokhsar said. “Even though there’s been an enormous amount of time, it’s still not long enough for them to move.”
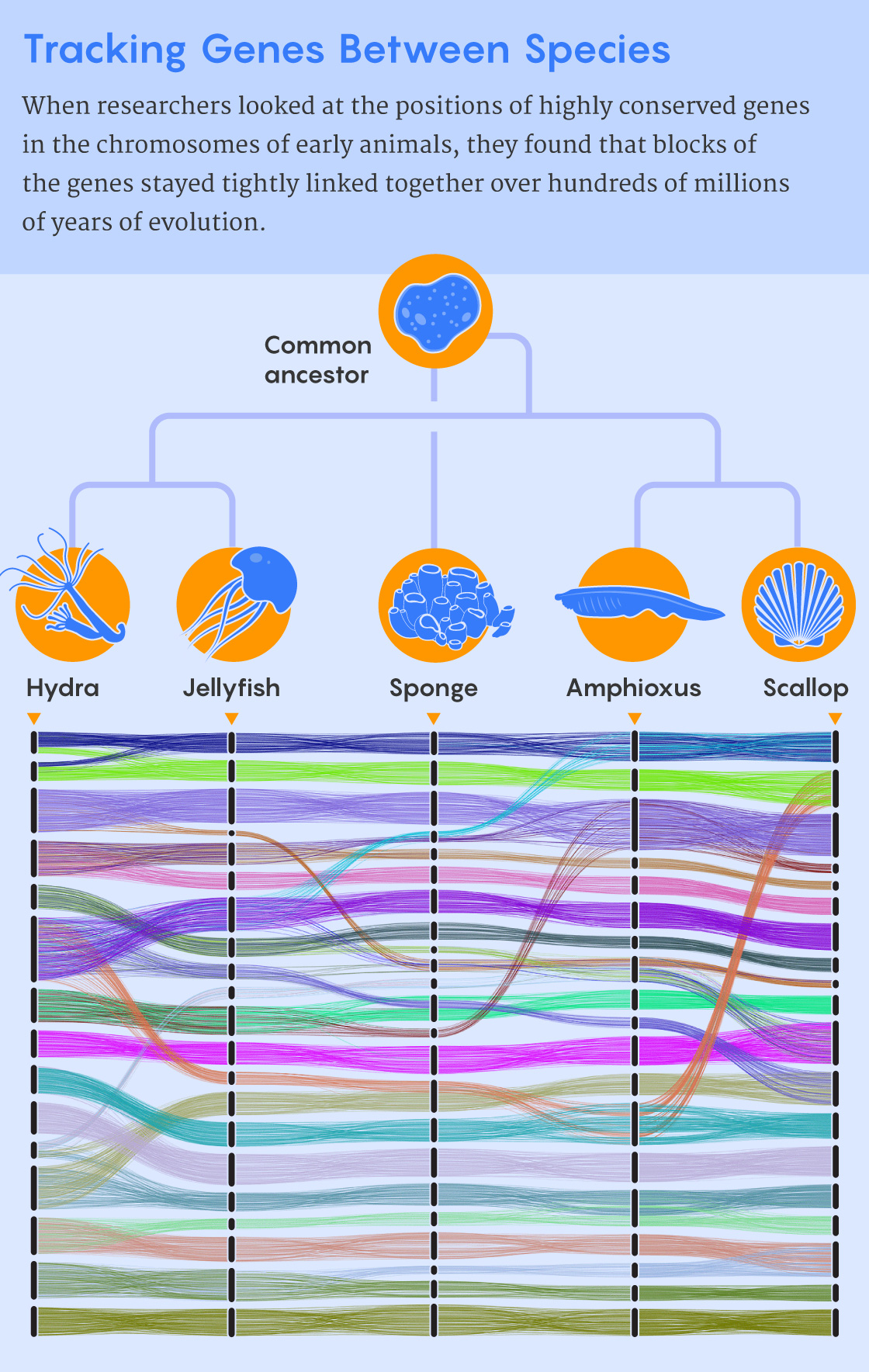
Samuel Velasco/Quanta Magazine; source: 10.1126/sciadv.abi5884
Rokhsar’s team found, however, that when those rare chromosome fusions did occur, they left a clear signature: After a fusion, the genes from the two blocks intermingled and were reordered because “inversion mutations” accumulated in them over time. As a result, genes from the two blocks blended like milk poured into a cup of tea, never to be separated out again. “There’s an entropic drive towards mixing that you can’t undo,” Rokhsar said.
And because the processes of gene block fusion, mixture and duplication are so rare, irreversible and specific, they are traceable: It’s highly unlikely for a chromosome to break at the same place twice and then fuse and mix with another gene block in the same way.
The signatures of these events in chromosomes therefore represent a novel set of derived characteristics that biologists can use to test hypotheses about how species are related. If two lineages share a mixture of two gene blocks, then the mixing most likely happened in their common ancestor. If lineages have two sets of the same gene blocks, a genome duplication probably occurred in their common ancestor. This makes the syntenies a “very, very powerful tool,”said Oleg Simakov, a genomicist at the University of Vienna and first author of the papers.
Fingerprints of Evolutionary Events
“One of my favorite aspects of our study is that we make predictions about what to expect in genomes that are yet to be sequenced,” Rokhsar wrote in an email to Quanta. For example, his team discovered that diverse invertebrates classified as spiralians all share four specific fusion-with-mixing patterns, which implies that the fusion events happened in their common ancestor. “It follows that all spiralians should show these fusion-with-mixing patterns,” Rokhsar wrote. “If even one spiralian is found that lacks these patterns, then the hypothesis can be rejected!”
He added, “You don’t get to make those kinds of grand pronouncements about evolutionary history very often.”
Samuel Velasco/Quanta Magazine; source: 10.1126/sciadv.abi5884
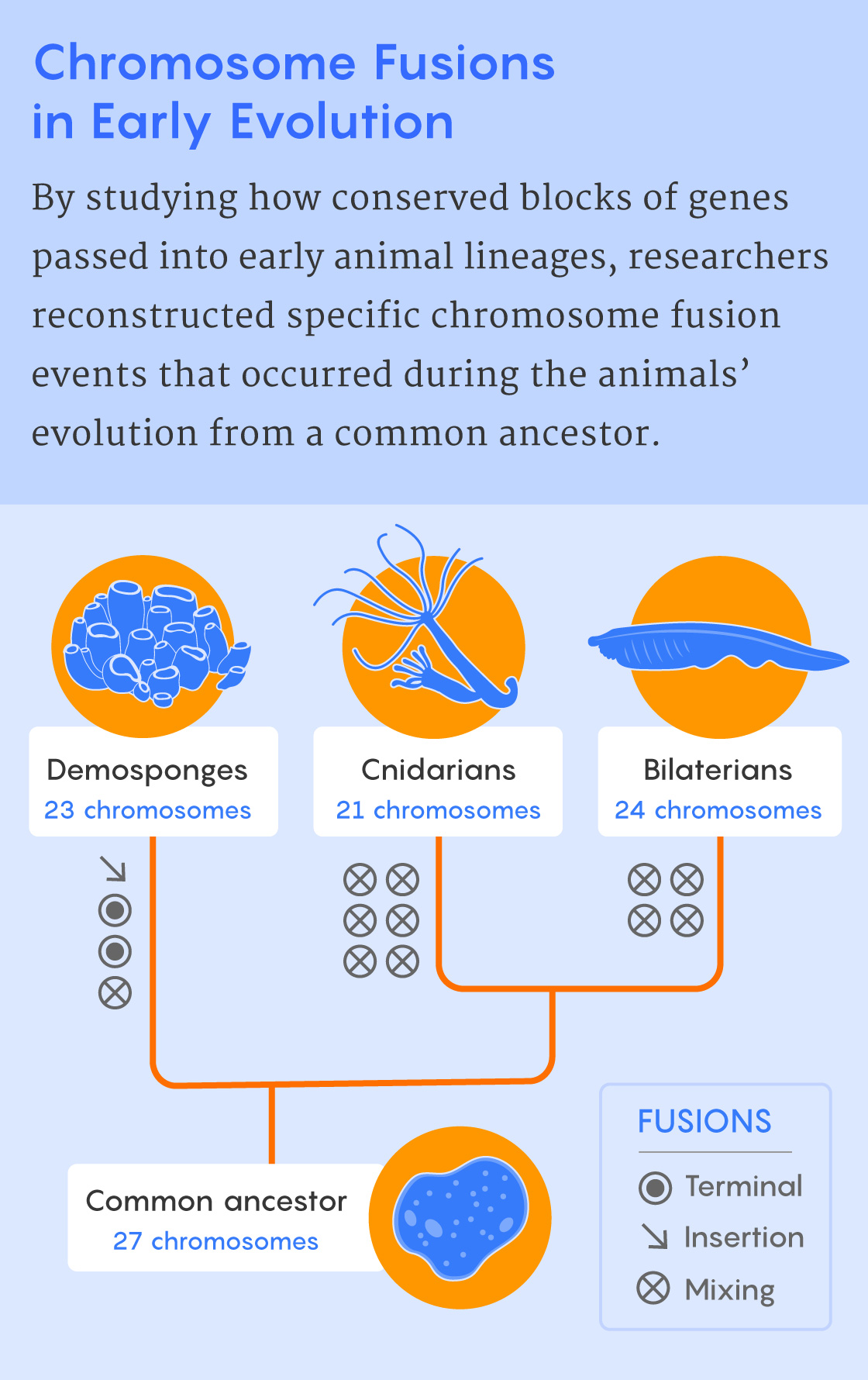
In their new Science Advances paper, Simakov, Rokhsar and their colleagues used the tectonic approach to learn more about the emergence of some of the earliest animal groups about 800 million years ago. Looking at the wide swath of animal life represented by sponges, cnidarians (such as hydras, jellyfish and corals) and bilaterians (animals with bilateral symmetry), the researchers found 29 blocks of genes that were highly conserved among their chromosomes.
Then, using the rules of chromosome fusing and gene mixing that they had identified, the researchers reconstructed the chromosome-level events that accompanied the evolution of these three lineages from a common ancestor. They showed that the chromosomes of sponges, cnidarians and bilaterians all represent distinctive ways of combining elements from the ancestral genome.
One provocative discovery was that some of the linked blocks of genes also seem to be present in the genomes of certain unicellular creatures like choanoflagellates, the closest relatives of multicellular animals. In multicellular animals, one of these blocks holds a diverse set of the homeobox genes that guide the development of their general body structure. This suggests that one of the very earliest events in the emergence of multicellular animals was the expansion and diversification of these important genes. “These ancient linkage units provide a framework for understanding gene and genome evolution in animals,” the scientists noted in their paper.
Their approach can distinguish between subtle but important differences in chromosomal events. For example, in their 2020 paper, the researchers deduced that the genome of vertebrates underwent a duplication sometime in the Cambrian period before evolution split the jawless and jawed fishes. They then found evidence that two jawed fishes hybridized later and underwent a second duplication of their genomes; this hybrid went on to become the ancestor of all bony fish.
John Postlethwait, a genomicist at the University of Oregon, emphasizes the significance of the team’s method of analysis. “They took a statistical approach, and they didn’t just say, ‘Well, it looks to me like such and such happened,’” he said. “This is a really important part of their methodology, not just that they had access to higher-quality genomes, but also that they took this quantitative approach and they actually tested these hypotheses.”
These studies mark only the beginning of what genome tectonics and genetic syntenies can teach us. In recent preprints shared on biorxiv.org, Rokhsar’s team reconstructed the evolution of frog chromosomes, and a European team looked at the chromosome evolution of teleost fish. A study in Current Biology found a “massive genome inversion” that drives the coexistence of divergent forms in common quails, hinting at some of the functional consequences of rearranging chromosomes.
It is tantalizing to hypothesize that the mixing of these gene linkage groups could be related to lineage diversification and evolutionary innovation during the last 500 million years. Chromosomal rearrangements may lead to mating incompatibilities that could cause a lineage to split in two. It’s also possible that a gene landing in a new neighborhood could have led to innovations in gene regulation. “Maybe this was one of the driving forces in the diversification of animals,” Simakov said.
“That’s the big question,” said Lewin. “These are truly tectonic shifts in the genome, and they are not likely to be without consequence.”