A Lack of COVID-19 Genomes Could Prolong the Pandemic
Introduction
Back at the beginning of the COVID-19 pandemic, before the disease had even drawn the attention of much of the world, researchers in China and Australia mapped the genome of the coronavirus isolated from one of the first patients in the Wuhan outbreak. This first genetic blueprint of the SARS-CoV-2 virus was publicly released soon after, on January 10, 2020. The disclosure of that genome, and others that soon followed, guided the vigorous international scientific response to the pandemic, including the timely development of diagnostic tests, surveillance strategies, vaccines and other new tools for managing the outbreak.
As a report by the World Health Organization (WHO) noted, because technology can now read the genome of a virus sample from a patient in just a few hours, “for the first time, genomic sequencing in real time has been able to inform the public health response to a pandemic.” All the successes that countries have had in coping with the pandemic are built on measures arising from our knowledge of its viral genome.
Yet the need to collect information about the SARS-CoV-2 genome is far from over. “It is too early to conclude how and when the pandemic will end,” said Meng Ling Moi, deputy director of the WHO Collaborating Center for Reference and Research on Tropical and Emerging Viral Diseases. Although numbers are dropping, more than 2.5 million new infections and more than 64,000 deaths were reported in the week before June 22, with significant increases in many countries. Health authorities attribute the strength and persistence of the pandemic to highly contagious variants of the virus that are spreading around the globe.
Genomic sequencing remains the foundational tool for understanding how the virus is evolving, and how our defenses against it need to adapt. In the hands of skilled researchers, genomic data can spell out the deepest secrets of the coronavirus, including epidemiological behaviors that patient data alone cannot capture.
Because of the singular importance of sequencing, in an online news conference last December, Tedros Adhanom Ghebreyesus, the director-general of the WHO, called on countries to step up their SARS-CoV-2 sequencing efforts. The European Commission followed suit a few weeks later, asking EU member states to sequence at least 5% — but preferably 10% — of positive test results from COVID-19 patients. The Centers for Disease Control and Prevention (CDC) also set a target of 5% for the United States in February.
The SARS-CoV-2 Sequencing Challenge: A Global Comparison
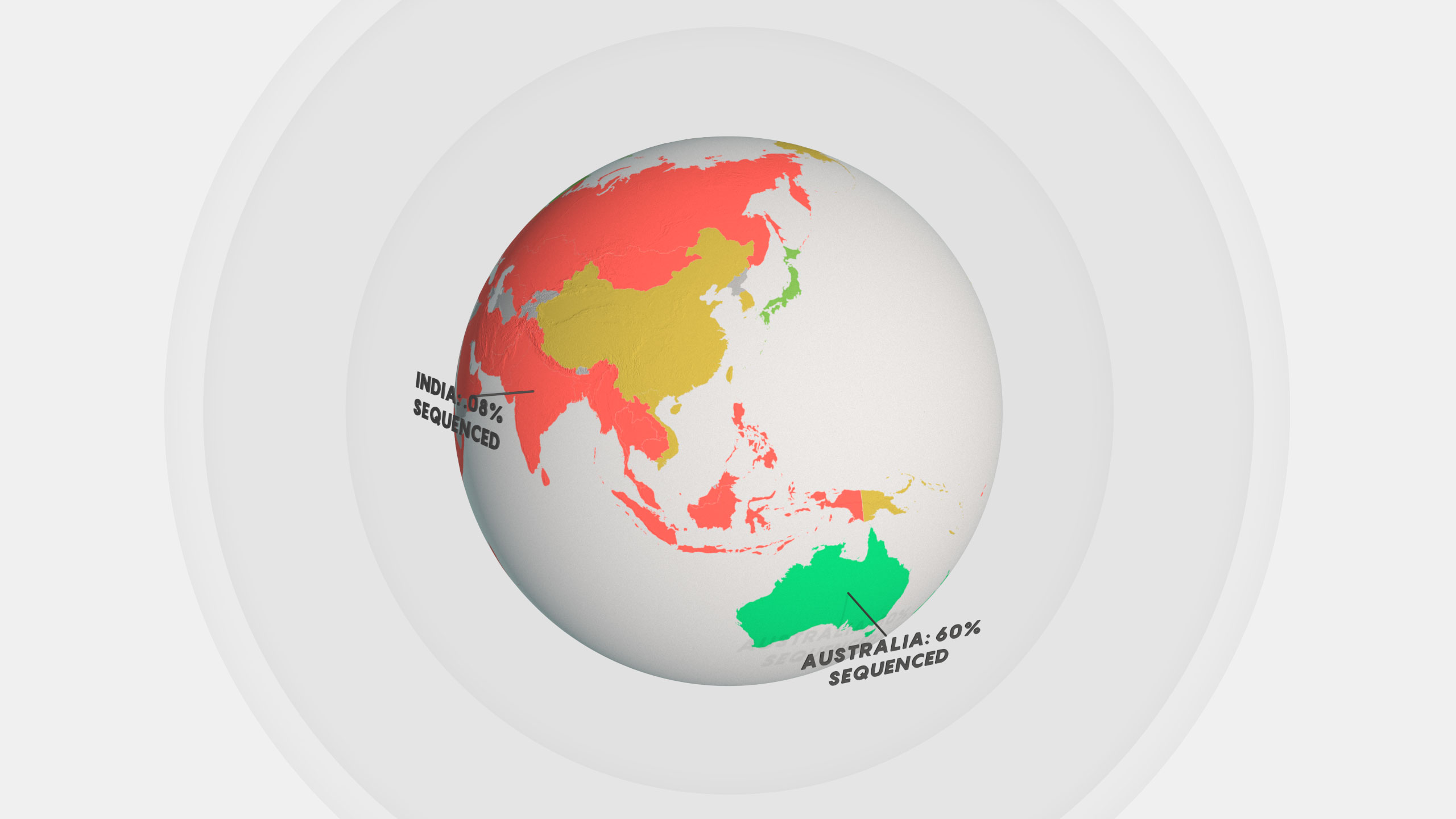
Global health authorities have recommended sequencing the viral genomes of 5% or more of all confirmed COVID-19 cases, but most countries are lagging far behind that goal. Choose a dataset for a map of the sequencing progress, caseload or number of genomes sequenced in each nation.
Percentage of COVID-19 genomes that have been sequenced
Number of confirmed COVID-19 cases
Number of viral genomes sequenced
Under 1%
10 million or more
0-999
1%-4.9%
1 million-9,999,999
1,000-9,999
5%-9.9%
100,000-999,999
10,000-99,999
10% or more
0-99,999
100,000 or more
To see the data for a specific nation, select from the drop-down menu or place your cursor over the map.
Yet countries are still falling far short of these goals, according to the Global Initiative on Sharing Avian Influenza Data (Gisaid), which manages the most widely used global repository of SARS-CoV-2 genomic data, and other sources. Almost 180 million confirmed COVID-19 infections have been reported globally, but only about 2,042,000 viral genome sequences — from barely more than 1% of the total cases — have been submitted. Rates of sequencing are improving in many countries, but not fast enough. The numbers are particularly alarming for the countries with the most infections: The U.S. has sequenced 1.7% of its 33.2 million cases, while Brazil, with almost 18.2 million infections, and India, with 30.1 million, have sequenced only about 0.1% of their caseloads.
Medical researchers worry that insufficient knowledge of how COVID-19 is changing could disastrously deepen and drag out the pandemic. “The more large gaps in our knowledge of the variants circulating globally, the more likely we are to miss the evolution of an important variant and find ourselves taking backward steps in the fight to control the pandemic,” said Justin O’Grady, who was until recently deputy director of the COVID-19 Genomics UK Consortium (COG-UK) and is now senior director of translational applications at Oxford Nanopore.
If the factors currently holding back viral surveillance are not addressed, there could be more uncontrolled viral outbreaks in our future. “If there are large ‘blind spots’ to virus sequencing surveillance, then with a fast-spreading infection, you cannot prevent pandemics,” said David Haussler, scientific director of the Genomics Institute of the University of California, Santa Cruz. “For this reason it is essential that virus genomes be sequenced everywhere in the world and that information be shared immediately.”
Why Viral Genomic Data Is Crucial
Scientists look at genome sequences collected throughout an epidemic as invaluable sources of clues about how a virus is evolving. The rate at which a viral genome mutates is relatively easy to estimate from laboratory work, but that isn’t the same as the virus’s rate of evolution, which depends on how quickly and successfully a mutation spreads throughout a population. The evolutionary rate is more accurately assessed by comparing the genomes of viral samples collected from different patients at different times. Studies suggest that for SARS-CoV-2, it is best for the samples to be collected at intervals of at least two months, with better results coming from more samples collected over longer periods.
Monitoring the evolution of SARS-CoV-2 is of course useful for developing and maintaining the accuracy of COVID-19 diagnostics: Testing could become unreliable over time or in different locations if the tests seek viral features that evolution has made obsolete. Vaccine design is also directly related to genomic information, particularly now that clinical medicine is using mRNA vaccines.
A visual representation of 56 SARS-CoV-2 genome sequences (left) reveals some of the differences in samples of the virus. Each genome is presented as a path (right) in which the colors signify the percentage of guanine and cytosine bases and the curvature signifies the proportion of repeats in each segment of the sequence.
A visual representation of 56 SARS-CoV-2 genome sequences (left) reveals some of the differences in samples of the virus. Each genome is presented as a path (right) in which the colors signify the percentage of guanine and cytosine bases and the curvature signifies the proportion of repeats in each segment of the sequence.
MARTIN KRZYWINSKI/Science Source
“We need to be able to make a vaccine for a disease before it is widely spread around the globe,” said Haussler. “With mRNA vaccines, we can get started as soon as the genome sequence of the infectious agent is available. This means that the most important issue now is getting genome sequences early in the outbreak of a new strain, before it has spread.”
Genomic information about SARS-CoV-2 has also uncovered the deep biology underlying COVID-19. It helped scientists identify the cellular receptor protein that SARS-CoV-2 binds to, for example, which in turn suggests which types of cells in the body are most vulnerable to infection.
But having a thorough record of the virus’s diversification throughout the pandemic is also important in a more subtle way because of what it can reveal about the disease’s epidemiology. Because the viral genome accumulates approximately one random change every two weeks, the number of genomic differences between samples can determine whether the tested COVID-19 patients are part of the same network of viral transmission.
“The virus’s genome acts as a molecular fingerprint,” said David O’Connor, a professor of pathology and laboratory medicine at the University of Wisconsin, Madison, whose laboratory analyzes SARS-CoV-2 genomes. “We were initially worried that health care workers were being infected by COVID patients in their care.” But the genomes of the viruses in the medics and the patients were rarely identical, “suggesting that the health care workers were infected in the community, not in the hospital,” he said.
Similar kinds of genomic epidemiology helped researchers determine very early in the pandemic that an outbreak in Connecticut came from multiple U.S. sources, and not directly from international travelers. It also helped researchers determine that in Brazil, non-pharmaceutical interventions like masks and social distancing decreased the spread of the disease within states but were less effective at slowing the spread between states.
Patterns of viral diversity in the population can be used to estimate important epidemiological parameters of COVID-19, such as its reproduction number, R0. They can also help to reveal dynamics of a pandemic that can’t be deduced from epidemiological data alone; for example, they can illuminate what happened in an outbreak before any cases were identified.
Analyses can also reveal subtle differences in the epidemiological behaviors of specific lineages or variants of the virus. Health authorities around the world are concerned about several significant variants of SARS-CoV-2 that spread more easily. Among them: B.1.1.7 (alpha), first identified in the U.K., which may carry a greater risk of death; B.1.351 (beta), first identified in South Africa, which may have extra resistance to some vaccines; B.1.617.2 (delta), which was first identified in India and has become a leading cause of new cases; P.1 (gamma), identified in Japan and Brazil; and B.1.427 and B.1.429 (both known as epsilon), first identified in California.
Samuel Velasco/Quanta Magazine; Source: CDC
More variants will undoubtedly emerge over time, and it is unclear how much these variants will complicate, or even set back, efforts to bring the pandemic to an end. “Ongoing genomic sequencing is key in identifying the emergence of ‘vaccine escape’ variants,” Moi said. This makes it all the more troubling that most nations have failed to even come close to the levels of genome sequencing that may be needed.
The state of the genomic surveillance situation is grimmest in 38 countries with reported COVID-19 infections but no sequencing data shared with Gisaid. These make up some of the poorest countries in the world, such as Chad and Burundi. The African continent, as of June 27, has reported more than 5.3 million infections (3.9 million of these are confirmed), but its countries have sequenced and released only about 22,700 genomes, or at best only 0.6% of its cases. More than 40% of those genome sequences (about 9,600) come from just one country, South Africa.
The consequences of the paucity of data on Africa could be serious for people everywhere. “Africa, given its human population variation, is a candidate to becoming the source of ever more pathogenic and refractory strains,” said Muntasar Ibrahim, a Sudanese geneticist and professor of molecular biology at the University of Khartoum, where he leads its Institute of Endemic Diseases.
Strategic and Structural Failures
Shortfalls in sequencing cannot be blamed simply on a lack of money. (Sequencing costs about $120 per SARS-CoV-2 genome, but the costs can be significantly lowered by sequencing the genomes in large batches, according to Haussler.) Some of the poorest countries have sequenced more of their cases than some of the richest countries, so wealth cannot be the only determining factor. Gambia, for instance, at 7.8%, has sequenced more than Germany (3.6%), a country with 60 times its gross domestic product per capita.
Nor do low rates merely reflect how hard countries have been hit by the pandemic. About 10% of the U.S. population has had COVID-19, resulting in a low sequencing rate (1.7%) even though the U.S. has sequenced the most SARS-CoV-2 genomes. But the U.K., where about 7% of the population has had the disease, has sequenced more than 10% of its caseload: It has only the 13th-highest rate of sequencing in the world, but it has sequenced more virus genomes than all the countries ahead of it put together.
What really seems to have determined the genome-sequencing performance of countries during the pandemic is a combination of their strategic choices and biomedical infrastructure.
Tom Maniatis, chief executive officer of the New York Genome Center (NYGC), noted that COVID-19 surveillance in the U.S. has been compromised by a systemic lack of connections between facilities that have samples of the virus — hospitals, public health laboratories and commercial testing facilities — and facilities with the capacity to sequence them. “Though the situation has improved, there have been persistent logistical challenges,” he said.
Maniatis and Soren Germer, who leads the sequencing and analytics teams at NYGC, said that obtaining samples had been the biggest challenge in the U.S. “During the early days of the pandemic when New York was particularly hard hit, even the most research-focused hospitals often did not have the resources to collect samples for research,” they explained by email. “We have heard stories of truly heroic efforts to save some of these samples for research and surveillance,” but the severely strained hospitals had to prioritize treating patients and protecting staff. Maniatis and Germer also pointed to a lack of coordinated funding in the U.S., which has been uneven at the state and local level and has only recently begun at the federal level.
Rolf Apweiler, director of the European Bioinformatics Institute, says that the nations depositing SARS-CoV-2 sequences into the dedicated genome data platform that his organization operates also vary substantially in their ambitions. While some countries aim low or have no genomic surveillance of SARS-CoV-2, he said, “countries like Denmark, Iceland, Australia and the U.K. aim to sequence between 10% of all positive samples in times of high infection rates and all positive samples technically feasible in times of low infection rates.”
The genome sequencing effort may already be bearing fruit for some of the countries engaging in it most vigorously. COG-UK is a consortium of genomic experts working to track, trace and control the SARS-CoV-2 virus in the U.K. It formed when the country’s scientists took steps early in the pandemic to ensure genomic sequencing at scale, aided by £20 million from the government. Within weeks of its formation in March 2020, the consortium had made the first sampled genomes publicly available; it has now sequenced more than 450,000 virus genomes.
O’Grady credits that work with helping to contain the pandemic in the U.K. “Genome sequencing identified the B.1.1.7 variant, providing us with an answer as to why case numbers were increasing dramatically towards the end of 2020 and enabling us to implement successful control measures,” he said. When other variants were discovered in South Africa and elsewhere, U.K. authorities increased the testing and contract tracing efforts and curtailed the spread of the variants into the country.
Preparing to Fight On
Many countries are now working to scale up their sequencing programs. In February, the CDC pledged $200 million as a “down payment” for genome surveillance. In April, the Biden administration dedicated $1.7 billion to boosting sequencing efforts and fighting variants of SARS-CoV-2. “The U.S. is now investing heavily in sequencing with the realization that the gains we’ve made are fragile and could be upended by viral variants,” O’Connor said.
In January, the Indian government set up the Indian SARS-CoV-2 Genomics Consortium to expedite the gene sequencing effort through a growing network of institutions. The nationally coordinated genome-sequencing program has sequenced more than 15,000 genomes in about three months, said Anurag Agrawal, a senior scientist with the consortium and director of the CSIR-Institute of Genomics and Integrative Biology in New Delhi, one of the participating institutions. “I expect the numbers to keep getting better,” he said.
The situation is improving in Africa, too. Segun Fatumo, an assistant professor of genetic epidemiology and bioinformatics at the Medical Research Council/Uganda Virus Research Institute, said that African governments urgently need to provide funding for relevant research and infrastructure. But he also noted that Africa has been moderately successful in the fight against the coronavirus, and genome sequencing has greatly contributed to this.
“The WHO has established a network of COVID-19 genomic sequencing laboratories across Africa” in 18 countries, he said. “Africa is central to human origin and disease susceptibility, so large-scale genomic study in populations of African descent might yield potential therapeutic strategies.”
Apweiler feels that a pandemic can be successfully managed only if it is tackled at a global level with as much coordination and collaboration as possible. “A problematic new lineage of SARS-CoV-2 in one country may become a worldwide problem very quickly,” he said. “Our response to the pandemic will be globally only as strong as the weakest part of the global efforts.”
Moi agrees about the importance of sequencing, but also suggests that it will always be necessary to balance that effort against other local priorities to ensure the best public health impact. “Particularly during large outbreaks, sequencing large numbers of virus [genomes] may not be practical” and could increase the burdens on laboratories and medical facilities that are already under pressure, she said. But she is also confident that “with optimal sequencing strategies in place, powerful insights can still be achieved with well-planned sampling and testing.”
Preventing Future Pandemics
“Had the pandemic happened even five years ago, it would have been a lot more difficult to implement genomic surveillance programs at scale,” O’Connor said. “The technologies to democratize sequencing and make it available to small laboratories and public health authorities simply weren’t available.”
The infrastructure and technology developed to map the virus could also be beneficial beyond COVID-19. “Our next hope is that the detailed observation of viral evolution during the pandemic and the research will help with the more rapid development of targeted therapeutics in future pandemics,” Maniatis said.
To him, the real question is whether the informational networks and infrastructure will enable viral surveillance to become routine, so that the discovery of the next potential pandemic virus can be a normal part of the public health system. The WHO has called the integration of genome sequencing into the regular practices of the global health community “a must” in preparations for future threats.
Haussler agreed that building global pathogen sequencing and genome sharing capability could help prevent future viral outbreaks. “It is one of the most important investments the world can make at this point,” he said. “It is likely to save many lives and many trillions of dollars in the long run.”