Cell by Cell, Scientists Map the Genetic Steps as Eggs Become Animals

A fertilized egg of a Xenopus frog goes through its first rounds of cell division in this time-lapse video. Scientists have now tracked how those cells find their developmental destiny by measuring which genes are expressed in each one during those first hours of life.
Olena Shmahalo/Quanta Magazine; Source: Enrique Amaya
Introduction
A fertilized egg divides first into two cells, then four, then eight and so on. Meanwhile, those cells progress from undifferentiated blobs in a cluster to more diverse identities associated with heart, brain, muscle, blood, bone and other tissues. Though the overall process is familiar, scientists have not understood it in much detail.
But three papers appearing today in Science are changing that, as they unveil work with major significance for the field of developmental biology. Using a combination of gene sequencing and mathematical methods, the researchers traced the patterns of gene expression in every cell in embryos of zebra fish and of Western clawed frogs through many stages of development during their first 24 hours.
The results revealed, at a previously impossible resolution and scale, the genetic and developmental trajectories that embryonic cells follow to their eventual fates in fully differentiated tissues. Surprising new insights emerged as well: Many biologists, for example, believed that embryonic cells always followed branching paths toward maturity that committed them irrevocably to certain fates. But the new data indicates that cells can, in effect, sometimes “loop back” to follow a different path, and that cells with different developmental histories can sometimes end up as the same type of cell.
The powerful techniques used in these reports, according to experts in the field, mark a new frontier in the ability to study development, cell fates and disease. “Whatever tissue you’re interested in studying, there’s something in this data set that should be of interest to you,” said Berthold Göttgens, a molecular biologist at the University of Cambridge who did not participate in the research but has been doing similar work in mouse embryos. Just as the rise of genome sequencing studies put biology on a different footing, he said, “this kind of foundational data will stand the test of time. It’ll be a landmark people will go back to.”
“There’s a whole universe of possibilities that data like this opens up,” said Alexander Schier, a cell biologist at Harvard University and an author on one of the studies. “Before, when we could only work with a few genes, or a few cells, or a few developmental stages, it was like we were seeing two or three stars. Now we can suddenly see an entire galaxy.”
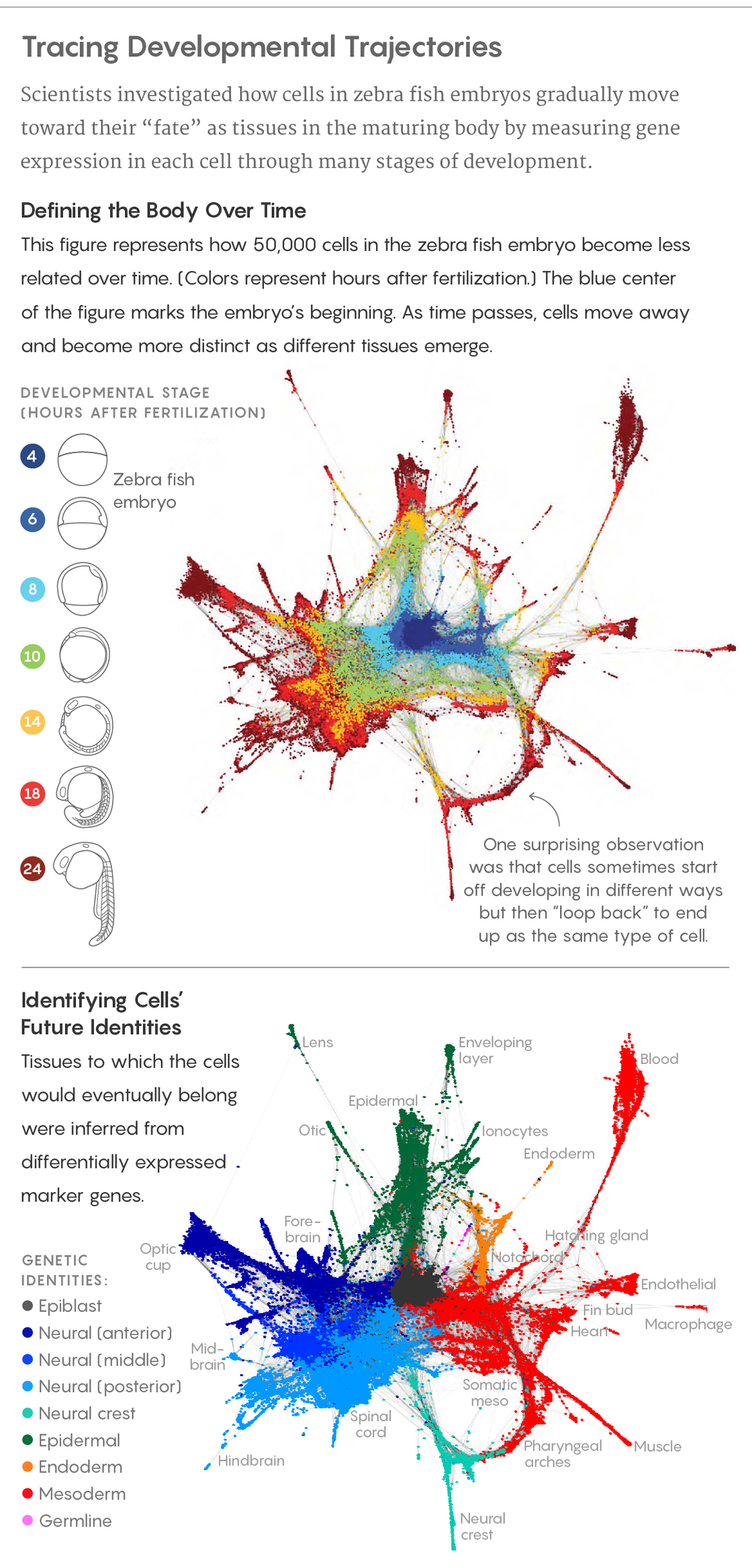
Adapted from D. E. Wagner et al., Science 10.1126/science.aar4362 (2018)
Traditionally, developmental biologists have dyed cells in a dividing embryo and tracked their spatial trajectory, or they’ve targeted specific genes and examined their effects on the organism. More recently, they’ve also used editing techniques to integrate “bar codes” into cellular DNA; as cells divide, the bar codes acquire mutations that can be used to determine shared cellular lineages.
The research published in Science takes a different approach. In their two papers, the Harvard systems biologists Allon Klein, Marc Kirschner, Sean Megason and their colleagues measured the expression of messenger RNA in each of the cells; one paper combined that data with the bar-coding technology. The messenger RNA data defined each cell’s identity based on which genes it expressed, while the bar coding provided information on where that cell came from — its “family history,” so to speak.
Analysis of that data confirmed many findings reported by other scientists after years of painstaking investigation. “It was exciting to see decades of developmental biology research encoded in our data,” Klein said.
But their experiments also pushed ahead to new findings, too. “With the old techniques, you could imagine getting [the equivalent of] a map at the city level. But you still didn’t really know what made, say, Philly Philly compared to Pittsburgh,” said Leonard Zon, a stem cell biologist at Harvard Medical School who was not involved in the studies. “Now you get a description of that character: how it’s put together, where it came from, what it’s doing.”
The process of development is often visualized as something like a tree, with the fertilized egg at its base and later cells branching out and specializing until all the cell types have emerged. And that’s certainly what it looks like at first: Schier’s group, which studied zebra fish embryos over their first 12 hours, confirmed this topology.
But at later stages, “thinking of development as a tree can be a poor description of what’s going on,” Klein said. In defiance of old assumptions, branches sometimes come together, converging onto the same path in a loop. Cells with very different developmental histories can end up reaching very similar destinations.
In short, cells may be more plastic than scientists thought, and may truly commit to becoming specific tissues relatively late. “It’s another way to think about how to make tissues,” Zon said.
The finding, Klein added, taps into a larger question about what’s really happening during the process of development, and the significance of all those intermediate states for cells that don’t end up in the adult organism. If the sole purpose of development were to create specific cell types, then theoretically, the needed genetic programs could be switched on in the cells right away: Mature muscles, bones, nerves and other tissues could be present in the embryos from almost the very beginning. But instead, development is a process, and the right cells need to emerge at the right times and places to lay down patterns for growth and transformation in the embryo. Cells in transitional states may have unique functions and developmental value in themselves, and not just as steps toward some end. Different parts of the embryo require different intermediate steps, even if they’re ultimately going to give rise to the same cell type.
That’s not the only kind of cellular plasticity that seems to be disrupting the tree metaphor. Schier’s group observed that certain cells expressed marker molecules earmarking them for more than one developmental fate: Even though they appeared to be going down one path already, the right outside signals could still prod them onto another. Schier speculates that these cells may be ones that live at the borders between tissues. Rather than following exact instructions to become one tissue type or another, they maintain ambiguity so that they can become either, depending on local interactions and cues.
The newly revealed plasticity of the developing cells can be dizzying, but the Harvard researchers also hope to find general patterns in their data that will reduce some of that complexity. Klein and his colleagues made a surprising discovery: Of the genes for proteins common to both the zebra fish and frogs they studied, only 30 percent were expressed in similar patterns with statistical significance. The animals expressed the rest in completely different ways, which suggested that the genes had adapted in their expression programs over the course of evolution. The conservation of the proteins at the sequence level seems to have no connection to the conservation of their expression in evolution.
“It came as a real shock,” Klein said. “It made us uncomfortable for a while.” But it also represented “another way evolution can tinker.”
He and his colleagues hope others will expand these comparative efforts to the rest of the tree of life. Göttgens, for instance, has been applying this approach to study both the formation of the blood system and the earliest diversification of cells that gives rise to the heart in mouse embryos. His work has focused on isolating genes that play key roles in those pathways — something Klein’s and Schier’s groups aim to do as well, by introducing mutations and observing how they affect cellular decision-making.
With even more data sets from a wider variety of organisms, researchers could also ask broader questions about how development has adapted throughout evolutionary history, and how cell types have evolved and emerged as a result. Take the differences that Klein and his teams observed between the gene expression levels in the zebra fish and the frog: If the scientists could find some rhyme or reason to explain which were conserved and which were not, it could help them lay down some ground rules about development. (Currently, Klein explains, it seems as though the genes with conserved expression patterns tend to be transcriptional regulators, which suggests “that evolution has laid down footprints for how to develop, while what it means to actually be a certain cell type has been subject to adaptation.”)
Scientists are excited about what this early embryo work could reveal about later stages of development, when cells within tissues and organs differentiate even further. Schier’s team is currently extending their work to the brain, for example, which comprises many more cell types than the 25 they mapped in the current study. That work might eventually provide important insights into how diseases develop. “If we don’t know, say, how all the different bits of the heart form, then it’s difficult to understand what’s actually going wrong in a congenital heart disease,” Göttgens said.
The benefits of that knowledge might extend to potential treatments as well. Klein’s and Schier’s technologies provide a recipe for how to make specific cell types, which could be used in regenerative medicine to manipulate cell fates deliberately. Knowing when to turn certain genes on or off — and accounting for cells’ plasticity and the importance of their intermediate stages — could go a long way toward helping bioengineers build better tissues. It could also help researchers improve cancer treatments. By better identifying the types of cells in a tumor, and their individual histories, researchers could better target the cells that are more malignant or drug-resistant.
In the meantime, though, the Harvard groups want to determine the mechanisms underlying the cell decisions they’ve described — to understand how different factors influence those pathways, why branching points occur where they do and which genes are responsible for those events. Such research, Klein hopes, will unmask some kind of global logic underlying their findings. Then, despite the complexity already seen, “we might understand that development is simpler than we think,” he said.
Correction: On June 13, the credit for the trio of time-lapse videos was updated to recognize the work of Huw Williams and James Smith.