Researchers Read the Sugary ‘Language’ on Cell Surfaces

Kevin Hong for Quanta Magazine
Introduction
When Pascal Gagneux envisions malaria parasites and other pathogens interacting with the surfaces of a host’s cells, he pictures a miniature rainforest with pathogenic particles flying overhead like colorful birds. The canopy consists of branching sugary molecules that adorn the surface of the cell. “If you’re a malaria parasite, you’re landing on a human red blood cell,” and “the first ‘leaves’ that you touch” are sugars called sialic acids, said Gagneux, an evolutionary biologist at the University of California, San Diego.
His ecological view of that interaction is rooted in his previous field work studying wild chimpanzee behavior in the dense West African forests. During those treks, he began to ask himself: “Why is it that humans and chimpanzees, who share so much of the same DNA, don’t deal with diseases the same way?”
“Diseases that give you and me the sniffles will actually kill chimpanzees,” he explained. But the opposite is also true. Chimps are not susceptible to influenza A, and HIV infections turn lethal in humans but stay mild in chimps. The malaria parasites that kill humans can’t infect chimps, and vice versa. This odd selectivity is not peculiar to primates — there are countless examples of pathogens devastating certain host species but not others.
Seeking an answer, Gagneux pivoted to the study of the glycomolecules, or glycans, in that “rainforest canopy” that shrouds cells. Glycans are a spectacularly diverse group of complex sugars (polysaccharides). They can exist on their own — cellulose is a plant glycan made up of long chains of glucose — or they can be anchored to other biomolecules like proteins and lipids, whose chemical properties they modify. Their structure can be linear (as in cellulose), but they can also be very highly branched, adding to their variety and complexity.
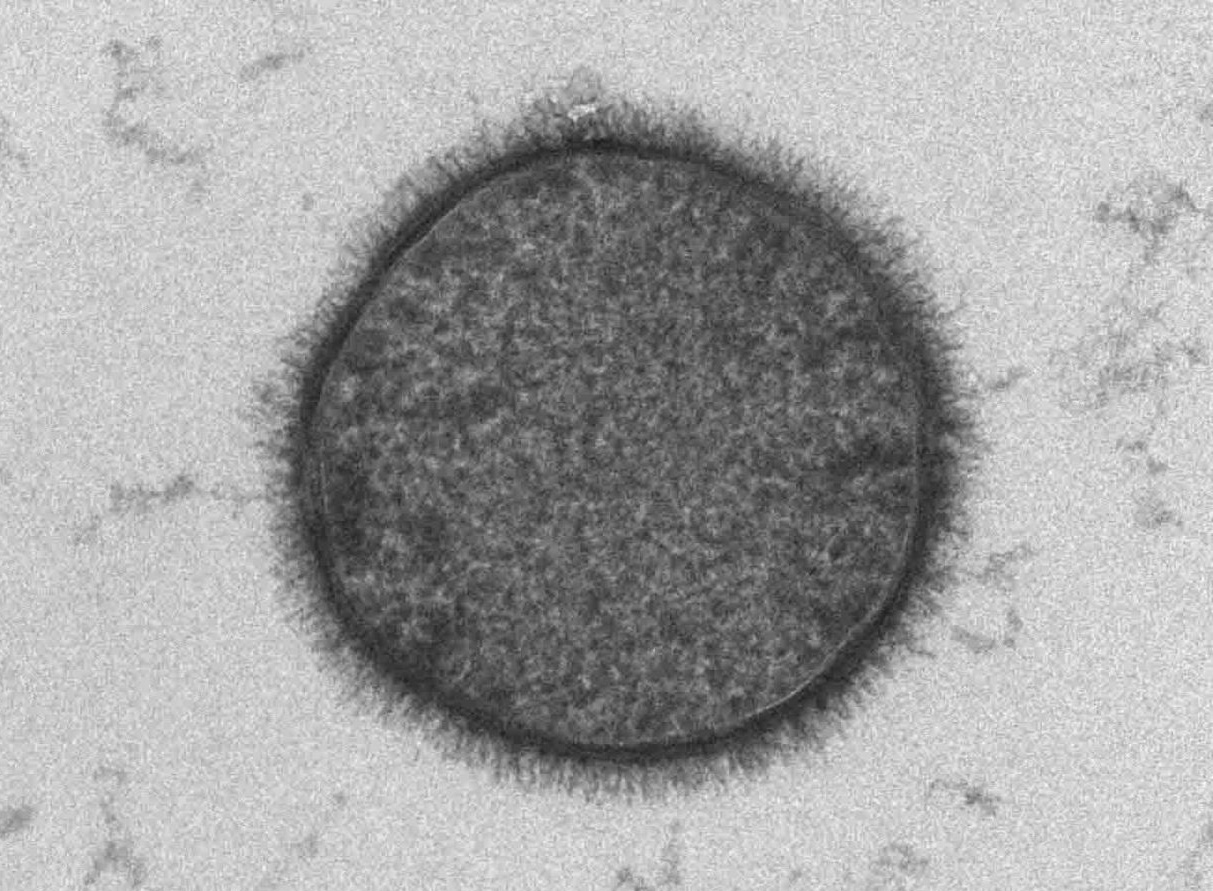
In this electron micrograph, glycans appear as a fuzzy coating over the surface of a bacterium.
Their endless variation among cells and species is central to why pathogens devastate certain host species but not others. It helps to explain the “spillover” of certain infectious agents, like SARS-CoV-2, from one species to another, leading to global pandemics. But it’s also a key to cellular behaviors even within species, such as the interactions between human sperm and the egg and uterine cells.
Now scientists may be verging on a breakthrough in the understanding of glycans and glycobiology. After analyzing a comprehensive data set of glycan structures and their known interactions, researchers at Harvard University and the Massachusetts Institute of Technology found a shared structural “language” that all organisms use when making glycans, like a municipal building code that ensures consistent, compatible architecture. The researchers have released a set of online tools that anyone can use to analyze glycan structures and functions.
Abundant but Mysterious
The shift in Gagneux’s interests happened when he met Ajit Varki, now a physician-scientist and co-director of UCSD’s Glycobiology Research and Training Center. Gagneux said that Varki, who became his mentor, had “just stumbled across the first biochemical difference between humans and chimpanzees.” Varki and his team had found that, more than 2 million years ago, a mutation in humans’ ancestors inactivated a gene that modifies sialic acids in all other primates and most other mammals. As a result, hundreds of millions of sialic acid glycans that are present in other primate cells are missing from human ones.
To Varki, glycans are still one of the greatest enigmas of the biological universe. They’re “actually so prominent, they’re a major component of biomass on the planet.” In fact, glycans make up most of the organic matter by mass: Cellulose and chitin, the major building material of arthropod exoskeletons and fungal cell walls, are nature’s two most abundant organic polymers. And yet in contrast with the overabundance of glycans, “this whole field has been left behind,” Varki said.
Daniel Bojar, a bioinformatics researcher at the University of Gothenburg and the Wallenberg Center for Molecular and Translational Medicine in Sweden, agrees that our knowledge of glycans pales in comparison to what we know about the other major biopolymers: DNA, RNA and proteins. Glycans, he explained in an email, “are a mysterious, omnipresent entity in biology that we either conveniently ignore or struggle to make sense of.”
According to Varki, the current state of glycobiology harks back to the late 20th century, when major changes were happening in biology. Glycans were heavily researched through the 1970s and the first half of the 1980s. “Glycans were very prominent, with one Nobel Prize every decade. There were very prominent people in many fields studying glycans,” he said.
But as Varki wrote in a 2017 review, “The field of glycosciences originated in ‘descriptive’ carbohydrate chemistry and biochemistry and remained in these domains for a long time,” instead of probing harder questions about the synthesis and functions of glycans.
Meanwhile, major technical advances were accelerating the study of nucleic acids and peptides, long linear molecules directly specified by genetic code templates. In contrast, the complex branching structures of glycans arise through a series of chemical reactions that add and modify sugar residues. There was no corresponding improvement in resources for studying them.
As a result, by the mid-1980s, “DNA, RNA and proteins, all the molecular biology, came and took off and left the glycans behind at the station,” Varki said. That development was disheartening for Varki, who was looking for his first independent research position around that time. But despite the challenges, he told himself, “I’m going to stick with studying these things,” even when many other researchers were giving up on them.
Samuel Velasco/Quanta Magazine; source: Dna 621
Gagneux said that “quite a lot of molecular biologists are borderline annoyed by glycans,” which are tiny and translucent. “You can only see them if you start throwing things at them that stick to them,” such as lectins, which are proteins that can tag short, distinctive saccharide sequences. Yet neglecting to study these critical components could mean missing game-changing information about some of humankind’s biggest challenges and questions.
Richard Cummings, a professor of surgery at Beth Israel Deaconess Medical Center and Harvard Medical School, describes his “life’s work” as focused on “understanding the structure of complex carbohydrates, these glycomolecules [and] how they’re made.” Glycomolecules, he said, are “the most complex structures that the human body makes.”
Cummings is a co-director of the worldwide Human Glycome Project. He and other researchers on that project, which was only launched in 2018, aim to “sequence and identify all of the glycans and carbohydrate structures — glycomolecules — in humans,” he noted. In contrast, the Human Genome Project launched in 1990 and formally concluded in 2003, illustrating just how big the gap has grown between knowledge of the human genome and the glycome.
Yet it is critical that researchers determine which roles specific glycans play in illness and disease if they hope to develop more effective strategies for preventing and treating these conditions.
Molecular Windows Into Disease
Some of that research is already proving fruitful. Huge strides have been made in the study of a growing group of rare genetic metabolic disorders stemming from defects in glycosylation, according to Varki. “After a slow start in the early 1990s an international effort of many investigators has now resulted in a veritable explosion in discoveries of human genetic disorders of glycosylation,” he wrote in his 2017 review article.
Researchers have already turned to glycomolecules to gain new insights about conditions as diverse as cystic fibrosis, cancers, sickle cell anemia, HIV and COVID-19. For instance, in 2020, Cummings and his colleagues published a Molecular Psychiatry review article covering 25 years of post-mortem brain studies on abnormal glycosylation in people with schizophrenia.
Cummings, who also directs the National Center for Functional Glycomics and the Harvard Medical School Center for Glycoscience, studies the function of glycomolecules in human biology and how mutations or alterations in those functions can cause pathologies. He also investigates how bacteria, parasitic worms and viruses such as influenza infect and sicken humans.
“It turns out in almost any of these cases, it is through interactions of glycomolecules that microorganisms and parasites cause human disease,” Cummings said. Linking that knowledge to new treatments or preventive measures often remains a grand challenge.
Decoding the Language of Glycans
One hurdle for glycobiology, Gagneux noted, is that even closely related species with high levels of genetic similarity, like chimps and humans, have glycans that can vary significantly because of constant, ongoing coevolution. Each species faces its own evolutionary pressures from diseases that leave a mark on its library of glycans: The host glycome evolves to evade or counter pathogens’ attacks, and the pathogens’ glycomes evolve to escape the immune defenses of their potential hosts.
“It gives rise to this molecular arms race that happens differently once you go separate evolutionary ways,” Gagneux said. For instance, even if you inject humans with chimp malaria parasites, they don’t get sick. (“Believe it or not, this was done [in Belgium] in the ’50’s,” he said.) That’s partly because the chimp malaria parasites can’t find the blend of sialic acids they seek on human red blood cells.
On the other hand, chimps are highly resistant to cholera because the Vibrio bacterium that causes the disease makes a toxin that targets only the sialic acids on the cells lining the human gut, punching holes through their membranes. Because of host-pathogen coevolution, “there’s a lot of diversity” in the glycome, Cummings said.
That diversity was apparent when scientists at MIT and the Wyss Institute for Biologically Inspired Engineering at Harvard used glycan-focused machine learning models to analyze a data set of more than 19,000 unique glycans. This included “6,969 eukaryotic, 6,119 prokaryotic, and 152 viral glycans,” they wrote in their 2020 Cell Host & Microbe study.
“Because we included all species for which we could find glycans, this dataset constituted a comprehensive snapshot of currently known species-specific glycans,” the researchers wrote.
Bojar, who was a postdoctoral fellow at the Wyss Institute and MIT at the time, is the study’s first author. He and his colleagues observed 1,027 unique simple sugars (monosaccharides) and chemical bonds in the glycan sequences. They treated these as “glycoletters” — “the smallest units of an alphabet for a glycan language,” they wrote. They then began looking through the data set for patterns of “glycowords,” defined as sequences five glycoletters long (that is, three monosaccharides linked by two bonds).
Samuel Velasco/Quanta Magazine
To that end, they trained a bidirectional recurrent neural network on sequences from their database and used it to create a model for a glycoletter-based language. Such neural networks are commonly used to learn and train language models. “You can kind of think about it as reading a sequence of text forward and then reading it backward,” said Rani Powers, a senior staff scientist at the Wyss Institute and a researcher on the study. “You want to keep the context of what is essentially the sentence in this case, rather than just pulling out all of the words or all of the letters out of context.”
In theory, the glycoletters in the data set could have formed nearly 1.2 trillion different glycowords. Yet, surprisingly, the researchers’ results indicated that only 19,866 distinct glycowords were present across all the available sequences. Notwithstanding the immense complexity and diversity of glycans, and the differences in glycans that are characteristic of various species, the evidence suggested that all organisms follow very similar rules in assembling them and use essentially the same biomolecular language to define their structure.
The researchers discovered that by fine-tuning their models, they could predict with high accuracy the taxonomic groups of the organisms from which glycans came. Furthermore, they were able to train the models to predict with about 92% accuracy whether glycan sequences in a reference data set were immunogenic to humans.
The results are “very exciting,” and the further application of sophisticated computational tools to understanding glycans could turn out to be “important and revelatory,” said Lara Mahal, a glycomics researcher at the University of Alberta who was not involved with the study. (She is working on a different project with Bojar.) “It helps reduce the complexity of glycans into clear patterns from which we can gather important information, for example on the pathogenicity of glycans,” she added.
The Wyss and MIT researchers hope that other teams will use the tools for glycomic design and analysis that they have developed and posted free online. According to Bojar, their most immediately useful application may be in the pharmaceutical industry, for glycoengineering therapeutic monoclonal antibodies. Antibody proteins latch onto specific antigen targets on pathogens. But it is the glycans linked to the proteins that determine how the antibodies interact with the rest of the body’s defenses and help to direct what kind of immune response follows. In the future, Bojar said, the tools might be able to suggest glycans that would improve the performance of antibodies, for example by limiting their side effects or more precisely calibrating their half-life in the body.
Mahal noted that she is already using the tools to learn more about the specificity of the assays used to identify the glycans on cells. “These new computational technologies combined with high-throughput analysis will revolutionize our understanding of the glycome and its role in disease,” she said.



