A Billion Years Before Sex, Ancient Cells Were Equipped for It

In researchers’ quest for the origins of the ability of gametes to fuse sexually, they are looking at related genes in primitive archaea and viruses.
Camille Chew for Quanta Magazine
Introduction
Most complex organisms engage in a strange bit of genomic math at some point in their lives: To multiply, they subtract and then add. That is, to reproduce through the process of meiosis they create specialized sex cells, or gametes, with half the usual number of chromosomes; they then merge pairs of those gametes to create new individuals with a full, unique genome. Sexual reproduction is nearly ubiquitous among eukaryotes — organisms from kelp to koalas that have a membrane-bound nucleus and organelles. We animals merge a sperm and an egg; mushrooms sprout from the underground collision of fungal threads; pollen sends tubes racing through floral tissues to join ovules, creating fruit and seeds where they meet.
Yet fusing cells like this runs contrary to the normal cellular life cycle. Cells divide into two through mitosis to reproduce asexually, but otherwise they mostly guard against major disruptions of their integrity, which could snuff out their lineage.
Why this hazardous arrangement caught on in evolution has been the subject of intense study; it’s generally presumed that sexual recombination helps to keep a species’ genome healthy and diverse enough to offset the risks. But researchers have also been piecing together molecular clues about how it evolved.
Now research recently shared on the biorxiv.org preprint server suggests that some cells had the power to fuse their membranes far back in their evolutionary history. The molecular machinery that makes this part of sexual reproduction possible may have existed more than 2 billion years ago in the simple prokaryotic cells called archaea, perhaps as much as a billion years before eukaryotes and sex evolved. But the new findings also hint at an explanation for why this kind of cell fusion for sexual reproduction didn’t appear earlier in life’s history, when it seemingly could have.
A Widespread Family of Fusion Proteins
Throughout the family tree of eukaryotes, the same protein enables the cell membranes and nuclear membranes of gametes to fuse: HAP2. Because it is shared so universally among eukaryotes, “the common ancestor must have had the same mechanism,” said Michael Brandeis, a cell biologist at the Hebrew University of Jerusalem in Israel.
But this last eukaryotic common ancestor may not have been the original bearer of the protein. HAP2 belongs to a superfamily of “fusexins,” which includes a class of proteins that some viruses use to fuse their viral envelopes with cell membranes. For evolutionary biologists, the relation of HAP2 to these viral fusexins poses a “virus or the egg” riddle. Did the first fusexins evolve in ancient eukaryotes, only to be pilfered by viruses for their own purposes? Or were fusexins a viral invention co-opted by eukaryotes at some point for gamete fusion? Finding the most ancient form of fusexin, the protein from which all eukaryotic and viral derivatives evolved, is critical to learning these answers.
Archaea were a natural place to search for answers. Because they are thought to be ancestors of eukaryotes — they were the original hosts of the symbiotic bacteria that became mitochondria and chloroplasts — archaea show what’s beneath the base of the eukaryotic family tree. Moreover, researchers have traced the origins of genes involved in the management and recombination of genetic material during meiosis to archaea. (Archaea appear to use these genes’ special abilities to repair other genes.) So archaea are where the international collaboration of scientists from Israel, Sweden, Argentina, Uruguay and other countries behind the new study began their search for answers.
Benjamin Podbilewicz of the Technion in Haifa, Israel, Luca Jovine of the Karolinska Institute in Sweden, Pablo Aguilar of the Institute of Physiology, Molecular Biology and Neurosciences in Argentina, and their colleagues predicted what a hypothetical fusexin in archaea might look like, based on the structures and protein sequences of HAP2 in modern eukaryotes. Further work, including scans of genomic databases, led them to 96 archaeal genes that seemed likely to produce proteins with shapes strikingly similar to the predicted fusexin structure, with some tweaks here and there. The researchers called these archaeal genes fsxA.
It seemed as though the team had discovered ancient fusexins in archaeal genomes, the first ever seen in that domain of life. But there was no guarantee that the proteins would behave like a fusexin in a living cellular system.
So Podbilewicz and his colleagues brought the whispered potential of those genes to life. They incorporated fsxA into cultured hamster kidney cells and enabled the expression of the genes. By modifying some of the cells to have fluorescent red nuclei and others to have fluorescent green nuclei, the team could check whether the cells were using the putative archaeal fusexins to fuse.
And that’s precisely what they saw. In the cells producing the FsxA protein, the green and red nuclei mixed five times as often as nuclei in unaltered cells did, because the cells with the fsxA gene were merging as if they were pairs of gametes.
The presence of functional fusexins in the archaea could mean that the molecular tools for sex predated the evolution of eukaryotes, the researchers suggest. Based on their evolutionary analysis of the genes, they think these archaeal fusexins are the most primitive form of the protein yet seen and are likely to be billions of years old.
Functions Before Fusion
Paulo Gonzalez Hofstatter, an evolutionary biologist at the University of São Paulo in Brazil who was not involved with this research, thinks that the findings fill a gap in our knowledge of evolutionary history. “But in some way, it is also a bit predictable,” he said, “because basically the whole machinery for meiosis has an archaea origin.”
It also might not be the final word on the origin of the fusexins. Hofstatter cautioned that it’s still possible for the most ancient fusexin to have been viral: The gene from the viral proteins could have passed into archaea and then into eukaryotes.
Whether or not the fusexins were truly an archaeal innovation, the new work raises important questions about the original function of these proteins.
Merrill Sherman for Quanta Magazine; source: DOI: 10.1371/journal.pbio.3000007.g001
Andrew Pomiankowski, an evolutionary geneticist at University College London, notes that while the FsxA proteins appear to allow cell fusion when expressed in a mammalian cell, it’s still unknown what these purported fusexins are doing in archaea. It’s harder to probe the inner workings of archaea than those of bacteria because archaea tend to be far more difficult to cultivate in laboratory settings, often requiring intense environmental conditions like extreme heat or pressure.
Nevertheless, he lauds the approach that Podbilewicz and his team have taken. “They’re raising exactly the right question: What the hell is this [fusion] gene doing in the prokaryotes?” he said.
One possibility is that the fusexins help archaea conduct a kind of genetic exchange that loosely parallels a sexual process. Hofstatter pointed out that some species of archaea perform a limited form of cell fusion and DNA recombination that vaguely resembles meiosis. “They just fuse, they recombine, and divide again,” he said.
A Replacement for Lateral Transfers
How fusexins passed from archaea into eukaryotes is also far from obvious. For Hofstatter, the most likely explanation is direct (or “vertical”) inheritance from archaea into the lineage that became the host cell in the archaea-bacteria symbiosis underpinning all eukaryotic life. Essentially, eukaryotes may have been born this way, repurposing old tricks from their forebears.
But there are reasons to be cautious about this explanation. Rachel Whitaker, a microbiologist at the University of Illinois, Urbana-Champaign, is surprised that the team found fusexins in a group of salt-loving archaea, the Halobacteria, but not in the Crenarchaeota group, which are thought to be more closely related to eukaryotes than Halobacteria are.
One potentially significant detail found by the researchers is that fsxA genes dwell within special sections of the archaean genome known as integrated mobile elements, which can move around within a genome and even transfer to genomes in other individuals, sometimes across species barriers.
If these fusexin genes are indeed a bit flighty, then it’s speculatively possible that the earliest eukaryotes took up these jumping fusion genes from an archaeal cell and later retooled the genes for a role in gamete fusion in an example of “exaptation,” the evolution of new uses for old traits. Fusexins that may once have been used for limited genetic transfer between archaeal cells may then have seeded the evolution of sexual reproduction — what the researchers jokingly called a “eukaryotic sexaptation” in their preprint. (Podbilewicz and his colleagues declined to be interviewed for this article, pending peer review for their research.)
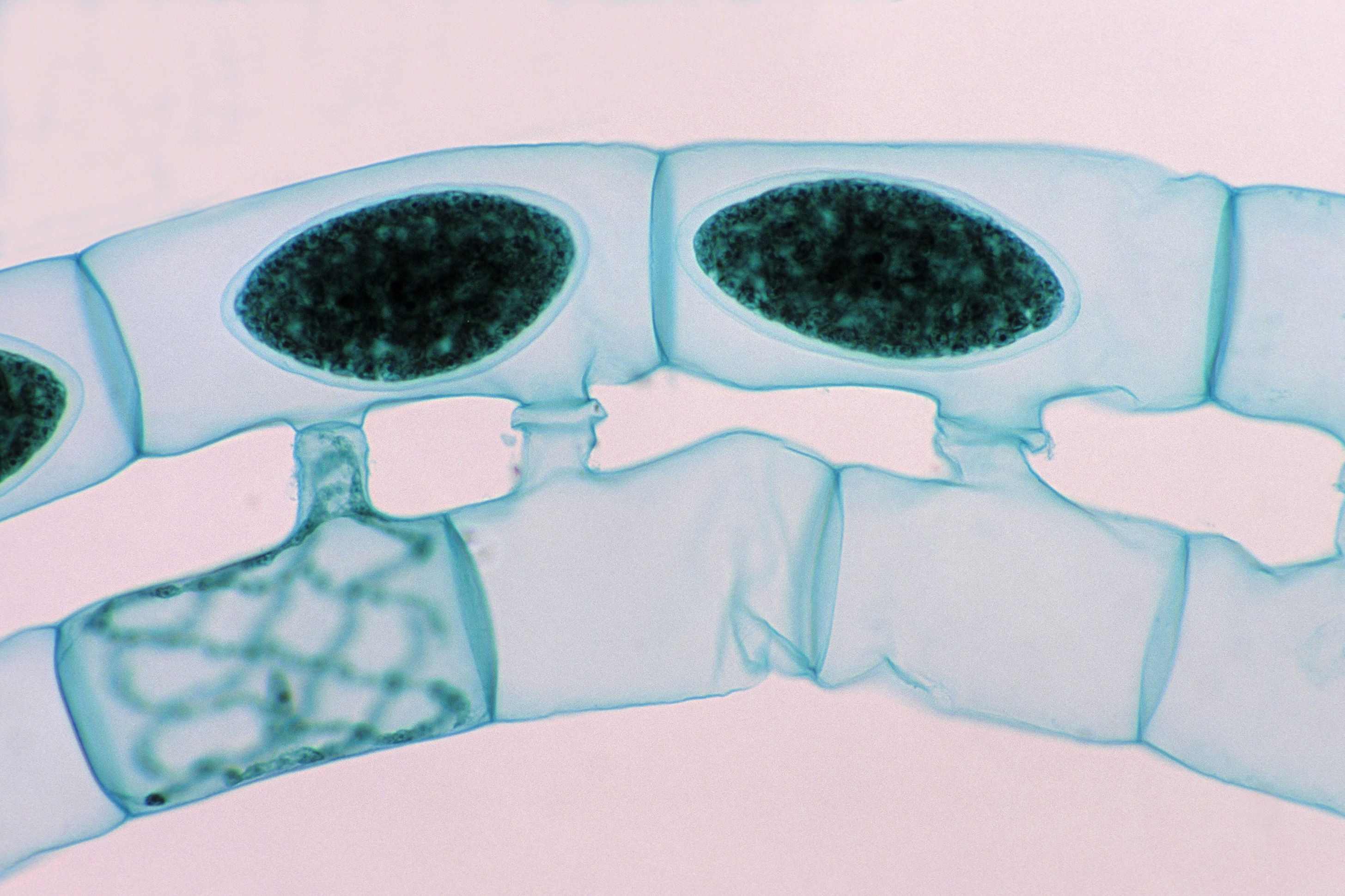
In the green alga Spirogyra, sexual reproduction occurs through the “conjugation” or fusion of cells in the filaments from different mating types. The dark ovals are the nuclei of the new zygotes produced through this fusion.
Ed Reschke
In any case, the discovery that ancient archaea probably had these fusion proteins has implications for our understanding of the evolution of eukaryotes. Indeed, it may offer clues about why sexual cellular fusion didn’t take off in prokaryotes even though the molecular underpinnings for it were present. The solution might be found in the ways that archaea collect new genes from one another and the environment, but also in the fact that the proto-fusexins reside in mobile genetic elements.
Pomiankowski explains that prokaryotes like archaea can exchange genes through viral infections or through plasmids, pieces of genetic material that can hop between cells. But they can also pick up DNA from the environment and recombine it into their genome, a process called transformation. These gene exchange practices may help maintain archaeal genomes, Pomiankowski said, by introducing new, potentially helpful genes and curtailing the accumulation of harmful gene variants.
But once archaea and bacteria permanently joined forces as eukaryotes, those cells became increasingly complex, and their genomes rapidly swelled in size. Consequently, the older “lateral gene transfer” strategy may have become untenable.
“The downside of lateral gene transfer is that it becomes less and less good if you have a bigger genome,” Pomiankowski said. “[If] you’re just picking a bit of DNA out of the environment, what’s the probability it’s going to match up? It’s a decreasing function of genome size.”
Cell fusion — made possible with repurposed ancestral fusexins — may have been instrumental in the transition to eukaryotes by allowing more coordinated, large-scale genetic exchange: sexual reproduction. Such a radical shift may have been more appropriate for maintaining the fledgling eukaryotic genome.
The Organisms With the Answers
Another possibility is that gamete fusion did not evolve primarily for the benefit of the eukaryotes themselves. Rather, it might have served the manipulative self-preservation purposes of ambitious genes. Because the archaeal fusexins are so closely associated with mobile genetic elements, Pomiankowski thinks fusexins could be “some kind of element that wanted to cause fusion between cells for its own benefit.” With the help of the integrated mobile element, the fusexin gene may have facilitated its own proliferation, first through prokaryotic fusion and then by eukaryotic fusion and genetic exchange.
But Pomiankowski acknowledges that this theory hinges on the integrated mobile elements in the archaea having been genuine jumper genes, which is uncertain. Such transposable elements are fairly uncommon in prokaryotes. For humans, “you’re 50% transposable elements,” Pomiankowski said. “But with bacteria, it’s like 0.5%.”
Cell fusion is an important part of sexual reproduction, but there were many other evolutionary steps between rudimentary fusing in archaea and the fusion of sperm and egg cells. Investigating this evolutionary transition may mean finding and studying organisms as close to the last shared ancestor of eukaryotes as possible.
Hofstatter thinks that will probably mean looking more closely at the single-celled eukaryotes called flagellates, which are highly diverse and abundant in aquatic habitats and are believed to be very similar to the last common ancestor of eukaryotes. “The key to understanding the origin of sex is the sampling of some flagellates that are not in culture so far,” he said.
Another place to look is the enigmatic Asgard lineage of archaea, which were described only a few years ago and are thought to be the closest relatives of eukaryotes. They may be able to provide some resolution to the question of how cell fusion and other processes came together to create sex, ushering in the vibrant, bustling tangle of eukaryotic life.