Math Reveals the Secrets of Cells’ Feedback Circuitry
Biologists have slowly made progress in understanding how cells and other living things implement negative feedback control systems. Finding the answers meant looking at the problem through an engineer’s eyes.
Peter Diamond for Quanta Magazine
Introduction
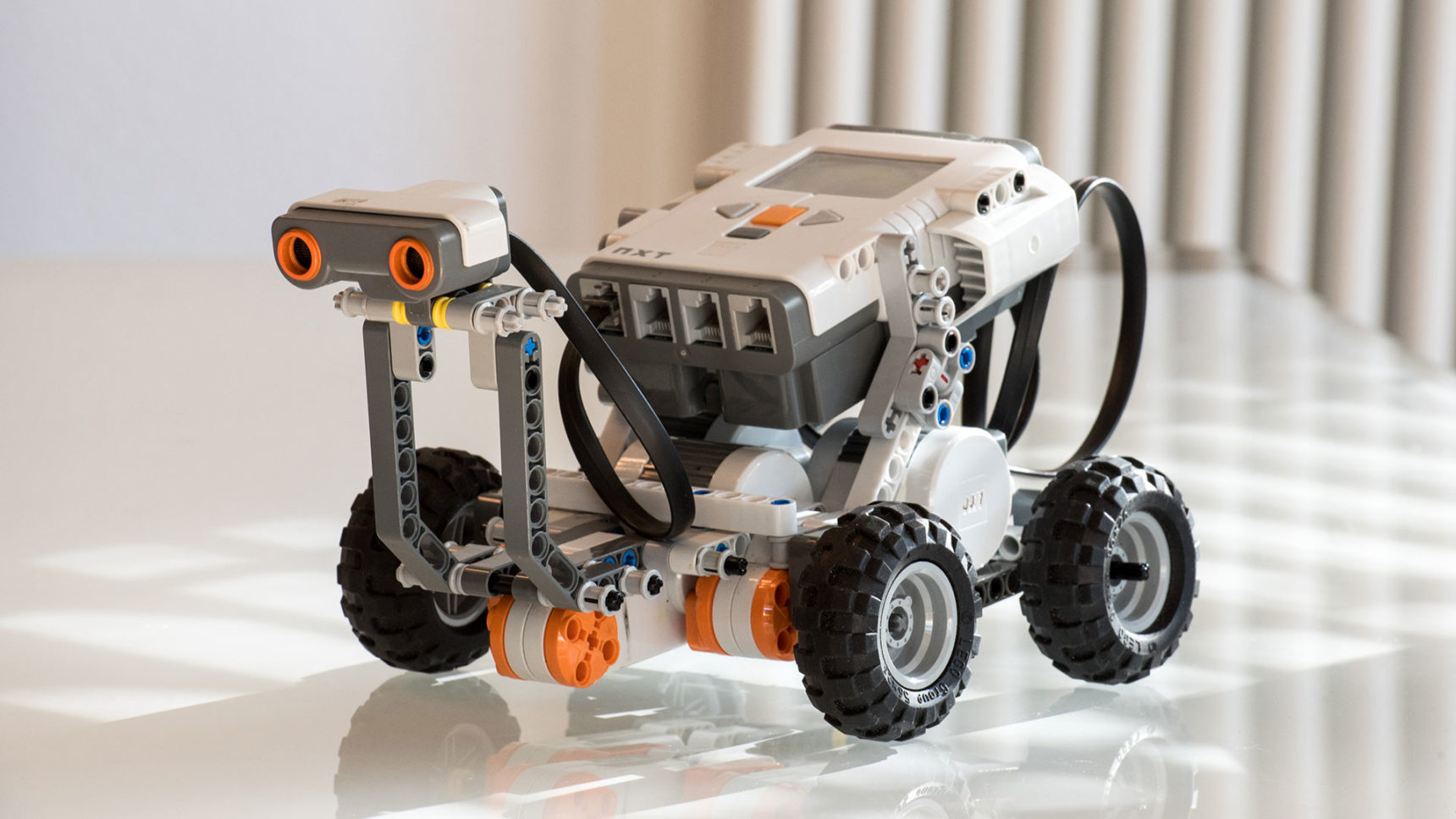
Mustafa Khammash’s small Lego robot is engaged in a one-way staring contest with a book held 30 centimeters in front of it. Khammash slides the book forward and his robot instantly revs its four offset wheels to follow it; he moves it closer and the robot leaps back, staying exactly 30 centimeters away from the book. Khammash weighs the machine down with his eyeglass case, he lifts the table at an angle, he replaces the wheels with ones that are 30% larger — each time, his robot restores its 30-centimeter buffer zone from the book and resumes staring at it.
The robot’s uncanny ability to correct its position gives it what biologists call robust perfect adaptation. “When the dust settles, there is no error,” said Khammash, a control theorist at the Swiss Federal Institute of Technology Zurich (ETH Zurich). “That’s the perfect adaptation; it keeps the distance perfectly.”
Whether in industrial control systems or in nature, negative feedback is an omnipresent strategy to help systems cope with disturbances. “People have noticed these feedback systems in physiology for as long as people have been studying physiology,” said Noah Olsman, a control theorist at Harvard University. Homeostasis, the self-regulation of biological systems, keeps many physiological parameters like body temperature, blood pressure and levels of blood glucose within exacting limits, whether we’ve run a marathon, gone scuba diving, or binge-watched Netflix all day. And for good reason: “If life couldn’t respond to changes and learn, it wouldn’t last very long,” Olsman said.
Vital as that negative feedback is, however, biologists have been hard pressed to explain how cells and more complex organisms implement feedback systems with the necessary responsiveness and precision. Only within the past couple of decades have they been able to sort out some of the fundamentals. Most recently, in an important advance this past summer, a team led by Khammash demonstrated a synthetic feedback system that could be installed in cells to help them adapt perfectly to disturbances, just like the robot. The work is backed by a mathematical proof that no simpler answer exists — a good indication that natural feedback systems probably work the same way.
Long before biologists figured out how nature accomplishes the feat, engineers were building feedback circuits into control systems to help keep airplanes on course, oil refineries pumping smoothly, and other automated systems humming along. (Control theorists call this set-point tracking with zero steady-state error.) Mathematically speaking, negative feedback can correct an error in three ways: proportionally, by considering the size of the error; integrally, by considering the amount of error incurred over the length of its duration; or derivatively, by considering how quickly or slowly the error is changing. The electronic proportional-integral-derivative (PID) controllers widely used in industrial control systems combine all three.
Of the three, integral feedback is the one that confers robust perfect adaptation; proportional and derivative feedback help mitigate disturbances but do not completely correct errors. The proof for this “is an old theorem in control theory,” said John Doyle, a mathematician at the California Institute of Technology. To work out how nature achieves robust perfect adaptation required a control theorist’s ability to spot a connection to integral feedback.
Negative feedback is a powerful example of the remarkable similarities between biology and engineering. In 1948, the mathematician Norbert Wiener proposed that regulatory systems in both animals and machines should be studied together, in a field he named cybernetics (from the Greek kubernētēs, meaning “steersman”).
“What math and engineering and biology have in common, at least modern engineering, is enormous hidden complexity,” Doyle said. Take, for example, a cellphone. It seems simple to operate, but underneath, many layers of control circuits are built atop one another.
“Biology’s kind of like that,” he said. “We live day to day in the complexities of our bodies; unless we’re sick, it’s largely automatic and unconscious. We are hardly aware of it.”
How Cows Integrate
An electrical engineer by training, Khammash first picked up a textbook on endocrinology at Iowa State University during the fall of 1998. His wife, who had just delivered their first child, had developed a postpartum thyroid disorder, and Khammash wanted to learn more about her illness. The text “could have been a book in control theory without the equations,” he said. “This hormone does this, this interaction increases the rate of that, and it closes the feedback loop, it’s the same story over and over again.”
Intrigued, Khammash went across campus to the National Animal Disease Center. There, he met the physiologist Jesse Goff, who suggested Khammash look into milk fever, a disease that older dairy cows get from severe calcium deficiency when they’re producing milk.
Calcium ions control many body functions, such as muscle contraction and neurotransmission. Blood calcium levels, kept within 8-10 milligrams per deciliter of blood, are therefore one of the most tightly regulated physiological variables in mammals. Milking drains calcium from a cow, creating a huge disturbance of blood calcium, Khammash says. And yet, in a healthy cow, blood calcium levels are always restored.
“As a control engineer, the first thing I thought is, ‘There’s got to be an integrator,’” he said. The question then became, “How do cows integrate?”
If a vehicle is going too fast or a robot is getting too close to an object, the vehicle’s driver can lift a foot off the accelerator and the robot can step back, directly reducing or reversing what has gone wrong. But in biology and chemistry, there is no subtraction: A protein’s concentration or a reaction’s rate cannot go negative. (Even if a cell stops making a protein, the existing molecules are still present.) Instead, everything has to be controlled with positives, the equivalent of a brake that opposes the influence of the accelerator. Some mechanism that performs mathematical integration is needed to calibrate how much pressure to apply to the brake and for how long.
To find an answer, Khammash enlisted the aid of his master’s student Hana El-Samad, who now leads her own research group at the University of California, San Francisco. They quickly ruled out the possibility that the integral controller comprised just one molecule; there had to be at least two. When that pair of molecules was eventually identified in 2002, they turned out to be well known to physiologists: the parathyroid hormone and a special form of vitamin D called 1,25-dihydroxycholecalciferol (1,25-DHCC).
When blood calcium plunges, the parathyroid gland produces more parathyroid hormone, which stimulates calcium ions to leave the skeleton and corrects the error proportionally. In turn, elevated parathyroid hormone levels ramp up the rate of 1,25-DHCC production in the gut, which promotes absorption of calcium in the small intestine. Because the rate of 1,25-DHCC production is tied to the concentration of parathyroid hormone, this feedback mechanism takes on a mathematically integral nature.
Khammash wasn’t the only one realizing nature uses integral feedback to achieve robust perfect adaptation. Earlier, in 2000, Doyle showed mathematically that the effectiveness of bacteria’s directed movements to find food was due to integral feedback. Later, El-Samad, Khammash and Doyle collaborated and showed that heat shock responses in bacteria — their production of protective “chaperone” molecules when overheated — are robust for the same reason.
Installing an Integrator in Cells
After solving the calcium problem, in 2002 Khammash and El-Samad moved to California. Khammash didn’t work on robust perfect adaptation again until he moved to ETH Zurich in 2011 and got the opportunity to start a synthetic biology lab. This time, the challenge was to artificially introduce a controller into cells. Such synthetic cellular controllers could one day help patients regain control over regulatory processes that have gone out of whack, such as insulin production in diabetics.
By this time, synthetic biologists had already built simple negative feedback circuits in cells that could correct errors proportionally. The first example, a rudimentary circuit in Escherichia coli, appeared in 2000. More recently, El-Samad reported installing a proportional feedback circuit with synthetic designer proteins developed by collaborators at the University of Washington. (This was significant because El-Samad showed that the designer proteins could be used in a modular fashion, akin to how most USB mice or printers nowadays are plug-and-play devices.)
Khammash set his mind on programming integral feedback into cells. “Any self-respecting controller should have an integrator,” he said, especially if it is to be robust.
But integral feedback is not easy to build. “You have to do it just in the right way,” Doyle said. Otherwise, the controller destabilizes. Instead of closing in on its target, an unstable controller will keep overshooting and start oscillating around it.
Khammash was joined in his efforts by Gabriele Lillacci, a theoretician then in the last year of his doctoral program, and Stephanie Aoki, a postdoctoral microbiologist. The trio moved into the BSA-1058 building in the Biopark Rosental in Basel and began setting up their new laboratory on the first floor. None of them had experience with synthetic biology.
The first design that Aoki and Lillacci tried was a simple circuit containing a pair of controller molecules: essentially, protein A, which turns on a gene for protein B, and protein B itself, which turns off the gene for A.
It didn’t work. Those were dark days for Aoki and Lillacci. “It doesn’t do what you’re expecting it to do, what you’re hoping it to do,” Aoki said. “You feel that you have no control over it.”
Part of the problem was that it’s just very difficult to engineer a cell. Translating well-established concepts in electrical and mechanical systems into biological terms is a big challenge, Olsman explained. “How do you actually take those ideas you can implement with resistors and capacitors, and implement them with proteins and RNA and DNA?”
And even when their E. coli cells finally did start to indicate that they could correct a disturbance, that result turned out to be an experimental artifact. “This was like one of the worst days, I think, in the lab for me,” Lillacci said.
The researchers didn’t realize it at the time, but their first design was inherently flawed. From a mathematical standpoint, unicellular organisms are very different beasts from whole creatures like cows: they are subject to statistical “noise.” Individual cells contain relatively few molecules, Khammash explained. Randomness from the probability that various molecules will meet, collide and react inside a cell comes into play much more forcefully.
Activators and Anti-Activators
On the eighth floor of BSA-1058, two theoreticians on Khammash’s team, Corentin Briat and Ankit Gupta, began discussing a new idea in early 2014. To minimize the effects of noise, the duo realized, the two controller molecules must have a very specific relationship: They have to bind to each other and neutralize each other’s biological activity. One must be the antithesis of the other.
On paper, Briat, Gupta, and Khammash came up with a new design. In this negative feedback loop, an activator molecule would stimulate production of a desired protein. In turn, that protein’s concentration would determine the rate of production for an anti-activator molecule, one that sequestered the activator. If something perturbed the system, any error in protein levels would be corrected through a corresponding change in the rate of anti-activator production. Best of all, because the activator and anti-activator molecules would seek out and neutralize each other, this loop would work even in a noisy cell.
Gupta proved mathematically that this design would result in a stable integrator for noisy cellular systems. But it was still completely theoretical. The trio had designed it without knowing what an antithetical pair of activator and anti-activator molecules would look like — or even if such pairs existed. And their unfamiliarity with the biology became an issue when a reviewer of their theory requested an example.
Khammash emailed a friend, the biologist Adam Arkin at the University of California, Berkeley, asking for help. Arkin quickly suggested the sigma and anti-sigma protein factors that are ubiquitous in bacteria. Arkin had previously used them to build a synthetic switch in cells.
Nor were sigma and anti-sigma factors the only options: There were also sense and anti-sense RNAs, and various toxin molecules and their antitoxins. “There’s tons of chemical reactions that do this,” Olsman said.
The theory was published in January 2016 to much excitement. “It was really clear how to implement this integration,” Olsman said. Two months earlier, Khammash had asked Aoki and Lillacci to drop the design they had been working on for three years and to try out the antithetical controller instead. “The theoretical foundation of it was just a lot more solid,” Lillacci said. The duo agreed to give the design a shot, using the same pair of sigma and anti-sigma factors Arkin had suggested.
It didn’t work — at least, not right away. Aoki and Lillacci had to navigate around two major assumptions that couldn’t hold in real life. One was that cells wouldn’t grow and dilute the factors involved. But they did, and with E. coli, cells doubled every 30 minutes or so. The other was that a gene’s rate of protein expression could be dialed up infinitely, whereas in reality, there is a limit to a gene’s expression.
In the fall of 2017, while his colleagues continued struggling in the lab, Gupta attended a conference in Ohio. There, he met other researchers also trying to build integrators in cells using the theory of the antithetical controller. All of them were struggling. Gupta thought there might be another design that would be easier to execute, making life easier for the experimentalists.
“It’s a legitimate question to ask whether, maybe, there is a simpler way,” Lillacci said. “And then it turns out that there isn’t.”
Gupta found that the mathematical constraints for robust perfect adaptation were so huge, they restricted the circuit designs that would be stable in a noisy setting. All of them required an antithetical pair of controller molecules.
Khammash and Gupta were elated at the mathematical proof that their approach, while arduous, was not just sound but inescapable. And for Aoki and Lillacci, who were already seeing signs that their cells could adapt to disturbances, the news kept them going.
“To find out that there’s really only one underlying topology that should be able to achieve this is really, really quite amazing to me,” Aoki said.
Finally, Aoki and Lillacci engineered a set of E. coli cells that were able to maintain a stable fluorescence, even in the face of disruptions from an introduced enzyme that ate up the green fluorescence protein. More dramatically, in another set of cells, when they lowered the temperature of incubation from 37 degrees Celsius to 30, the cells maintained their growth rate. Both Gupta’s proof and Aoki and Lillacci’s experiments were reported in Nature this June.
Olsman hopes this example will help bring a more rational, math-based approach into synthetic biology, as in engineering. “We don’t build a thousand airplanes, put them in the sky and hope they don’t fall down,” he said.
And beyond robust perfect adaptation, many other cryptic biological phenomena await deciphering, Doyle hopes, with the help of mathematics.