New Cell Atlases Reveal Untold Variety in the Brain and Beyond
Peter Greenwood for Quanta Magazine
Introduction
In the 16th century, the Belgian cartographer Abraham Ortelius created the world’s first modern atlas — a collection of maps that he called “The Theater of the World.” The maps, drawn by Ortelius and others, detailed what was at the time the best knowledge of the world’s continents, cities, mountains, rivers, lakes and oceans and helped usher in a new understanding of global geography.
Similarly, the creation of cell atlases — maps of organs and bodies constructed cell by cell — is heralding a new era in our understanding of biology. Powerful sequencing and imaging technologies invented in the last decade are revealing with unprecedented detail the composition of human organs and tissues, from the pancreas and liver to the placenta, as well as those of other animals like the mouse and fruit fly. With these new tools, researchers can fingerprint individual cells based on which genes they are expressing. That information has revealed subtle and unsuspected distinctions among cells and has begun to illuminate how the diversity of cell types can be essential to the healthy functioning of organs.
“We’re at this amazing point in time in science where we’re now able to understand the composition of these cell types,” said Steve Quake, a bioengineer and biophysicist at Stanford University who helped develop the technologies that make cell atlases possible. “It’s changed the way we understand how human biology works.”
Two cell atlas efforts, part of the National Institutes of Health’s $250 million brain cell census, that just released their findings illustrate the excitement bubbling up in the field. Today in Nature, a coalition of laboratories published nine studies that collectively form a detailed atlas of the mouse brain — the most comprehensive mammalian brain atlas to date. It describes more than 5,300 types of cells found throughout the organ. How these cells are distributed and are related to one another suggests many intriguing ideas about the evolution of the mammalian brain.
Last month, in another landmark contribution to the cell atlas project, a global network of collaborators published a suite of papers in Science journals that documented the incredible cellular diversity and complexity of the human brain. It is not a complete atlas because the scientists profiled only samples of cells across the organ. Nevertheless, it offers the most detailed map of the human brain ever made.
Because of that work, it’s clear that the human brain contains at least 3,300 distinct types of neurons, each expressing a different set of genes, which hints that the cells could serve a range of slightly different functions. Much of this diversity, it turns out, is in the less studied and more evolutionarily ancient regions of the brain.
This human brain cell atlas is merely an early edition. As scientists continue the task of tallying and classifying every single cell in the brain, they will likely identify thousands of additional cell types.
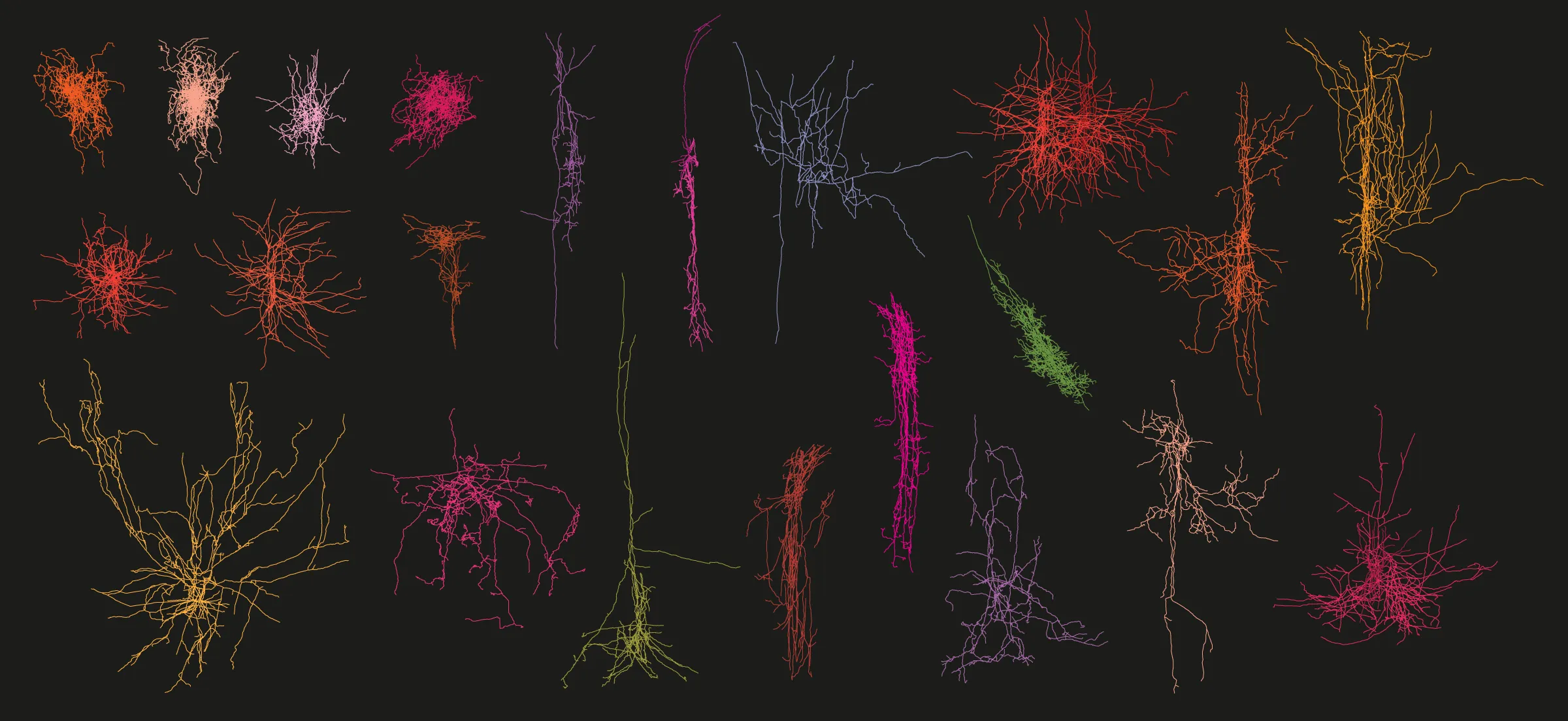
The human brain cell atlas is not complete, yet it has already documented more than 3,300 different types of neurons — a selection of which are portrayed here. Researchers expect to discover thousands of additional human brain cell types.
Neurons by Brian R. Lee and Rachel Dalley; Composition by Quanta Magazine
Locating and describing the variety of cells is only the first step. Scientists then have to work out what functions the cells serve and how they fit together. For that reason, Quake and others are working to stitch together cell maps from across the human body into a full human cell atlas: a holistic reference map of a human being that could help to tease apart which specific cells in which tissues go wrong to cause various diseases. The first draft of that atlas is likely to be published around two years from now, according to researchers familiar with the state of the work.
Although it’s still early days, scientists predict that these maps will answer fundamental questions about cells, how they function, and how they can change over an organism’s life span. “In a way, it’s a little bit of a leap of faith, almost the same way that the Human Genome Project was,” said Michael Angelo, an associate professor of pathology at Stanford. However, “it’s a very justified leap of faith.”
A Cell’s Unique Expression
Until just a few years ago, based on analyses of our tissues under microscopes, most researchers believed that the approximately 36 trillion cells in an adult human body could be categorized into only a few hundred distinct types: three types of muscle cells, epithelial and fibroblast cells in the skin, various kinds of neurons in the nervous system, endothelial cells lining the blood vessels, and so on. But the successes of the Human Genome Project invited researchers to look at our cellular makeup more deeply, at a genetic level.
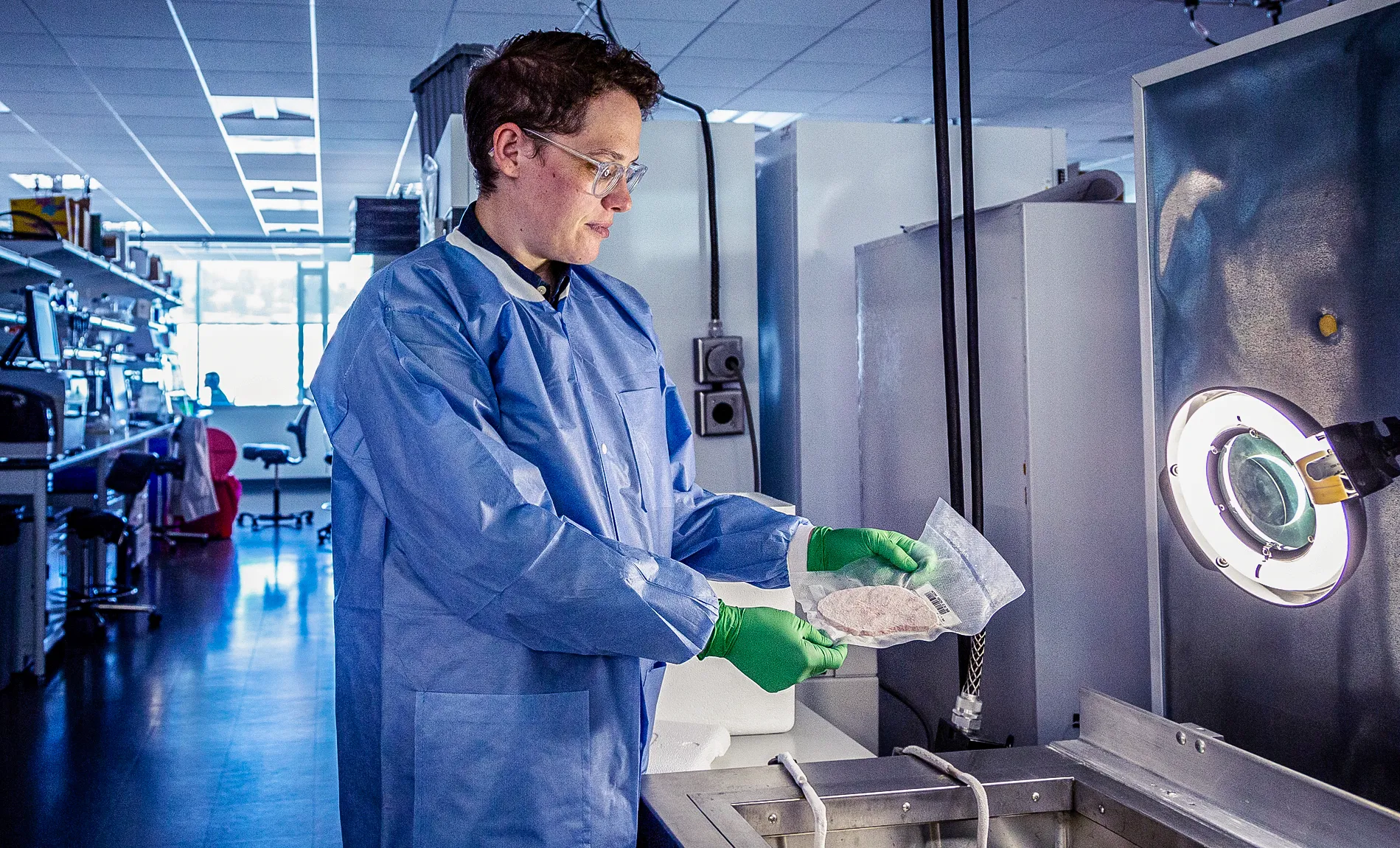
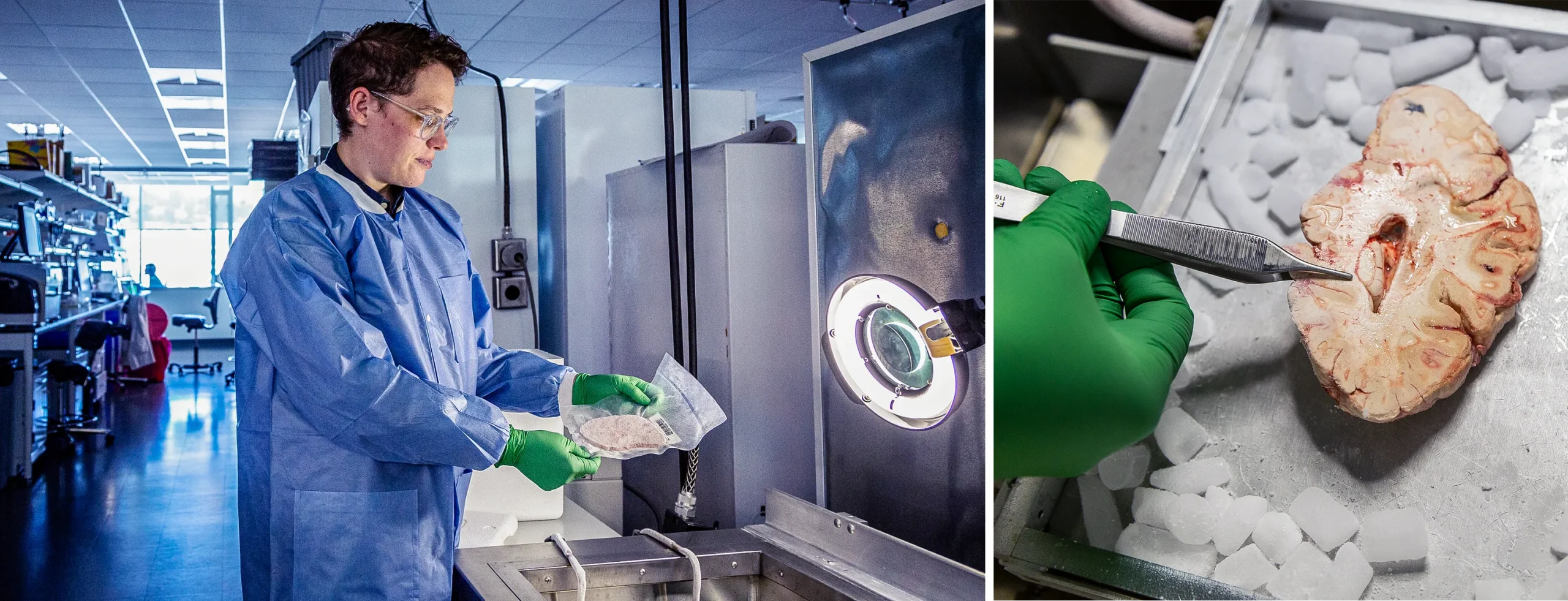


To build the human brain cell atlas, Rebecca Hodge from the Allen Institute had to carefully extract genetic data from slices of donated human brains.
Eric Dinnel/Allen Institute
The differences in our cells don’t come from differences in the genes themselves: Every cell in your body contains a copy of your distinctive DNA and the 20,000 genes for making proteins it encodes. “The genome is a parts list,” Quake said. “There’s no way to predict which cell types come out of a given genome.”
To distinguish a white blood cell from a muscle cell at the molecular level, scientists have to look at the cell’s RNA. RNA transcripts that are copied from DNA sequences ferry the instructions for building proteins to the ribosome, the cell’s protein construction center. Therefore, RNA transcripts contain the information about which genes are active and expressed in a particular cell.
Using a powerful set of single-cell genomics tools, scientists can read these expression patterns to fingerprint a cell. These tools have matured over the past few years so that scientists can now quickly and efficiently look at tens of thousands or even millions of cells in a single experiment.
“Once that was in place, then there’s nothing stopping you from making an atlas,” Quake said. “It all just kind of cascaded.”
Cell atlases started pouring in. Scientists began by examining the gene expression in individual cells from model organisms like fruit flies and mice. Since then they’ve moved on to humans. Within the last year, maps of blood vessels, tumors, placenta, kidney and intestine, among other tissues, were published in high-profile journals. Many of these atlases not only looked at cell identities, but also mapped out precisely where those cells were located in the tissues — information critical for understanding which cells might be involved with disease.
“The amount of data collection is mind-blowing,” said Elizabeth Rhea, a research assistant professor at the University of Washington who is not involved in any cell atlas effort. She reflected on how rapidly the technologies have advanced: 12 years ago, as a graduate student at Vanderbilt University, Rhea would examine the expression of a single gene in the developing mouse brain and liver. Now “we are able to do that, but for every gene expressed in every cell within a chunk of tissue,” she said.
With each new cell atlas, researchers have had to reckon with how much complexity they were missing before. Reviewing the data sometimes made Angelo feel as if he were looking at endlessly looping videos of fractals, he said. “You keep zooming in” and the pattern goes on indefinitely; the more you uncover, the more you realize there’s more to uncover. “A lot of this is kind of daunting.”
This has proved true for many human organs — especially the brain.
Mind-Blowing Diversity
The complexity of the human brain, in its construction and function, has limited our ability to understand it. Its 86 billion neurons are tiny sparks animating thoughts, perceptions, feelings and important functions throughout the body. Fabian Theis, director of the computational health center at Helmholtz Munich, who works on several atlas efforts but was not involved in the brain atlas, remembers one colleague telling him that the brain is like a separate organism. “It’s like 100 organs meshed into one,” he said.

Claudia Doege of Columbia University discovered that the hypothalamus, an ancient brain region, contains immense cellular diversity.
Courtesy of Claudia Doege
Claudia Doege, an associate professor of pathology and cell biology at Columbia University, echoed that view. “You need to put a lot of people together to dig through that big mess,” she said.
Doege studies the hypothalamus, an ancient and critical part of the brain that evolved to control basic bodily functions like body temperature, heart rate and hunger. In 2020, she was invited to help interpret brain atlas data collected from hypothalamic tissues — a rare opportunity to probe the infrequently sampled brain region.
It was a “gargantuan task,” said Hannah Glover, a postdoctoral researcher in Doege’s lab. Intermittently cursing at her computer, Glover spent months tearing through literature and reference atlases to identify genes expressed in different regions of the brain and comparing them to the genes found in the newly analyzed hypothalamus.
The analysis revealed “an extreme amount of diversity,” Doege said. They identified more than 350 distinct neuron populations and 19 non-neuron cell populations from 10 different regions within the hypothalamus.
Other research groups working on the brain cell census backed this up. Rebecca Hodge, an assistant investigator at the Allen Institute for Brain Science in Seattle who co-authored several of the new Science papers, also found that most of the cell types were located in evolutionarily ancient parts of the brain.
“That was one of the first descriptions of so much cellular diversity being outside of the cortex,” Hodge said, referring to the much-studied brain region where higher-order functions like critical thinking occur. “It’s scratching the surface, probably, of what the actual cellular diversity is there.”

Hannah Glover of Columbia University spent many months analyzing data from the hypothalamus to fingerprint cell types, each of which express a different set of genes.
Maria Caterina De Rosa
Across the entire brain, the research teams discovered more than 3,300 different types of neurons — and that account is surely incomplete. The most comprehensive atlas of the mouse brain to date, published today, identified 5,300 cell types, including both neurons and glia. The project is missing cell census data from some areas, such as the medulla. Still, researchers think that the full count of cell types for the human brain, when it is compiled, will likely be similar.
“We think that there are probably not too many more cell types in the human brain compared to the mouse brain,” said Hongkui Zeng, the director of the Allen Institute for Brain Science, who led the mouse brain cell atlas effort. The researchers have already compared brain atlas data sets from the two species, and based on what she’s seen, Zeng puts her personal estimate at “probably somewhere between 5,000 to 10,000” cell types in the human brain.
The mouse brain work does more than enumerate the cell types: It also places these cells in the context of their brain region as a navigable atlas. From that added positional information, the researchers drew fascinating new conclusions.
They found that, as in the human brain, the more ancient regions of the mouse brain, such as the hypothalamus and amygdala, contain most of its cellular diversity. However, these various cell types are closely related to one another and have relatively few differences. In contrast, the more evolutionarily novel brain regions, such as the cerebellum and thalamus, contain fewer cell types overall. But their cell types are highly distinct from one another, and each brain region contains cell types unique to it.
The authors hypothesized that because the ancient hypothalamus carries out basic functions critical for supporting life, like metabolism, breathing and reproduction, it has been evolutionarily constrained. It had longer to evolve, resulting in more cell types, but the cells couldn’t drift too far from their basic type. “The circuits are well established, and every time you make a change, you may jeopardize the animal’s life,” Zeng said.
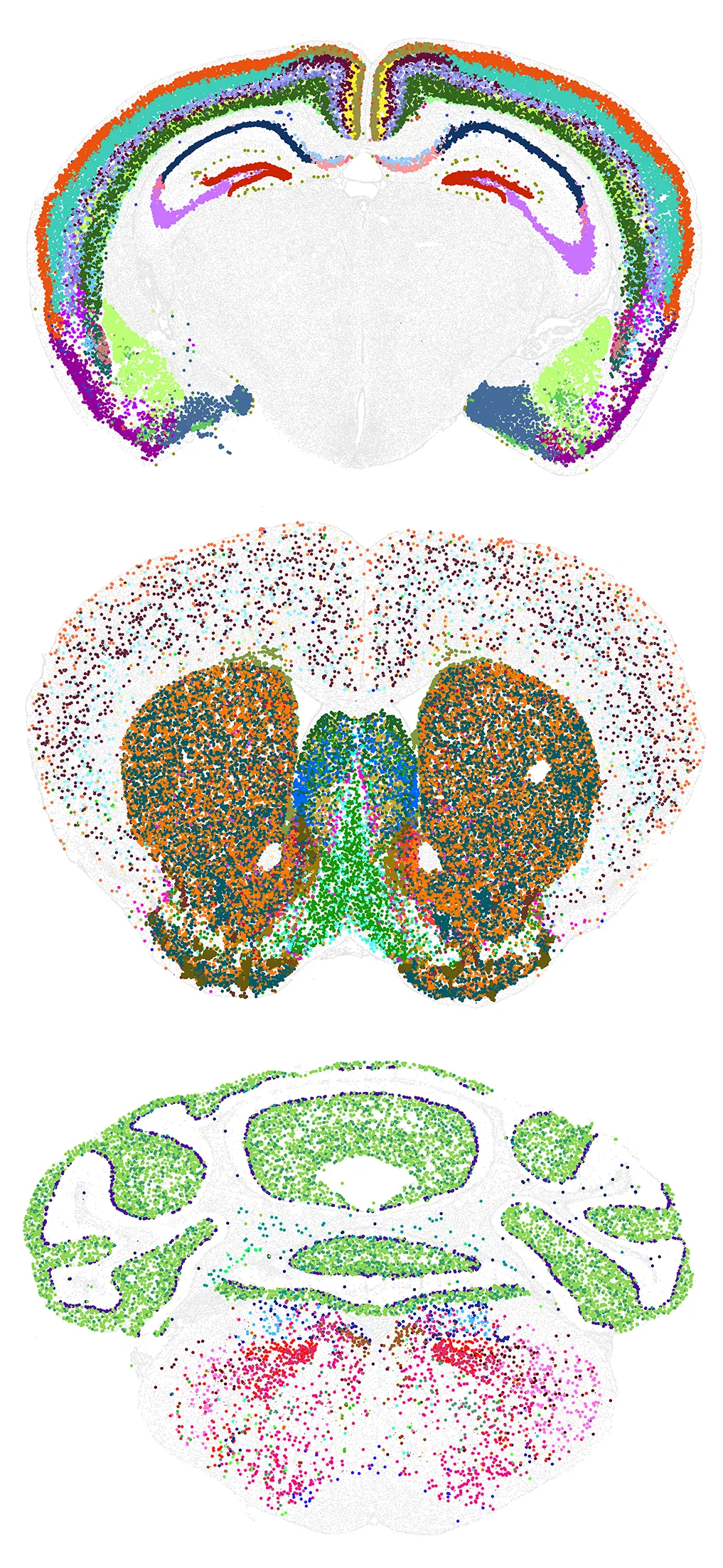
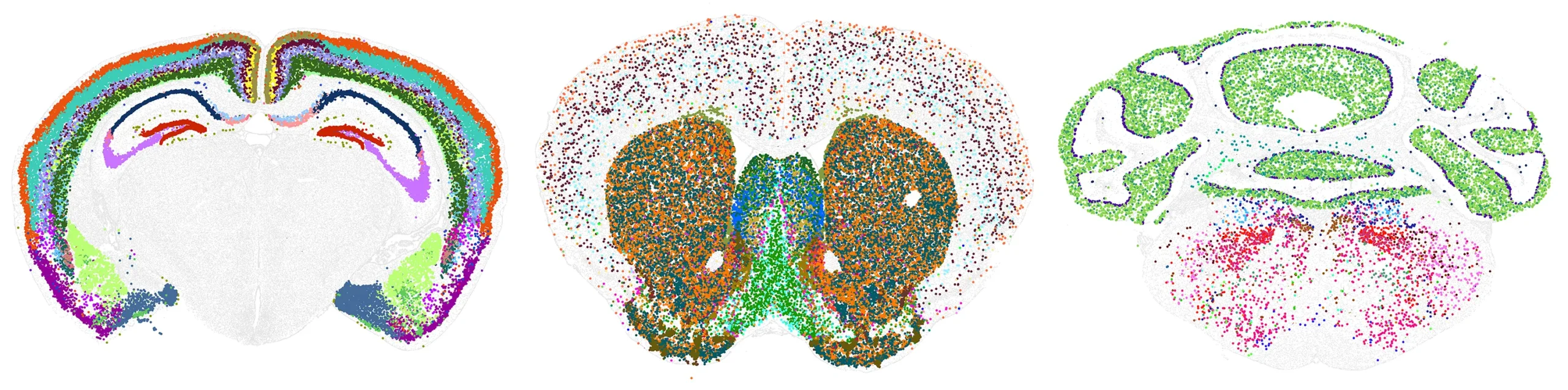
The new mouse brain cell atlas is the most comprehensive map of a mammalian brain ever created. In these images, each color represents a different cell type. The researchers discovered that more evolutionarily novel brain regions contain fewer types of cells, but those that are present are highly unique.
Courtesy of Allen Institute; source: https://www.nature.com/articles/s41586-023-06812-z
The newer brain regions like the cortex, on the other hand, carry out functions like cognition and emotion that help animals adapt to new environments and challenges, she said. These less constrained brain regions have been able to evolve faster and further, resulting in the brain cell types distinctive to the regions.
The findings from the mouse brain preview the kinds of discoveries scientists are likely to make when the human brain atlas is completed. Already, the human work is providing “basic facts about brain evolution that no one knew before,” said Thomas Naselaris, an associate professor at the University of Minnesota who isn’t involved with any atlas efforts. “No one has any idea what these new facts imply for the evolution of human cognition. But they are very basic facts that no one knew, so they are probably going to turn out to be important.”
Eventually, researchers plan to use the cellular detail of these new brain maps as guides for improving treatment of neurological disorders. “One of the things that I think is critical about making these atlases is that it provides a baseline or reference for understanding how things are changing in the context of disease,” Hodge said.
For example, despite decades of research, scientists still don’t really know what causes Alzheimer’s disease. It’s possible that our understanding of the disorder is held back by an oversimplified map of the brain’s composition. The specificity of a cell atlas could make research and medications more precise and impactful.
A Bridge to the Full Body
The findings inside the brain cell atlas alone are dizzying. But then consider that other cell atlases are uncovering similar trends. In organ after organ, researchers seem to be finding a greater diversity of cell types than they expected. These discoveries suggest that the healthy functioning of tissues often depends on having the right mix of cells doing slightly different jobs. Each individual cell atlas therefore provides almost incalculable new possibilities for understanding a region of the body, and all the things that can go wrong with it.
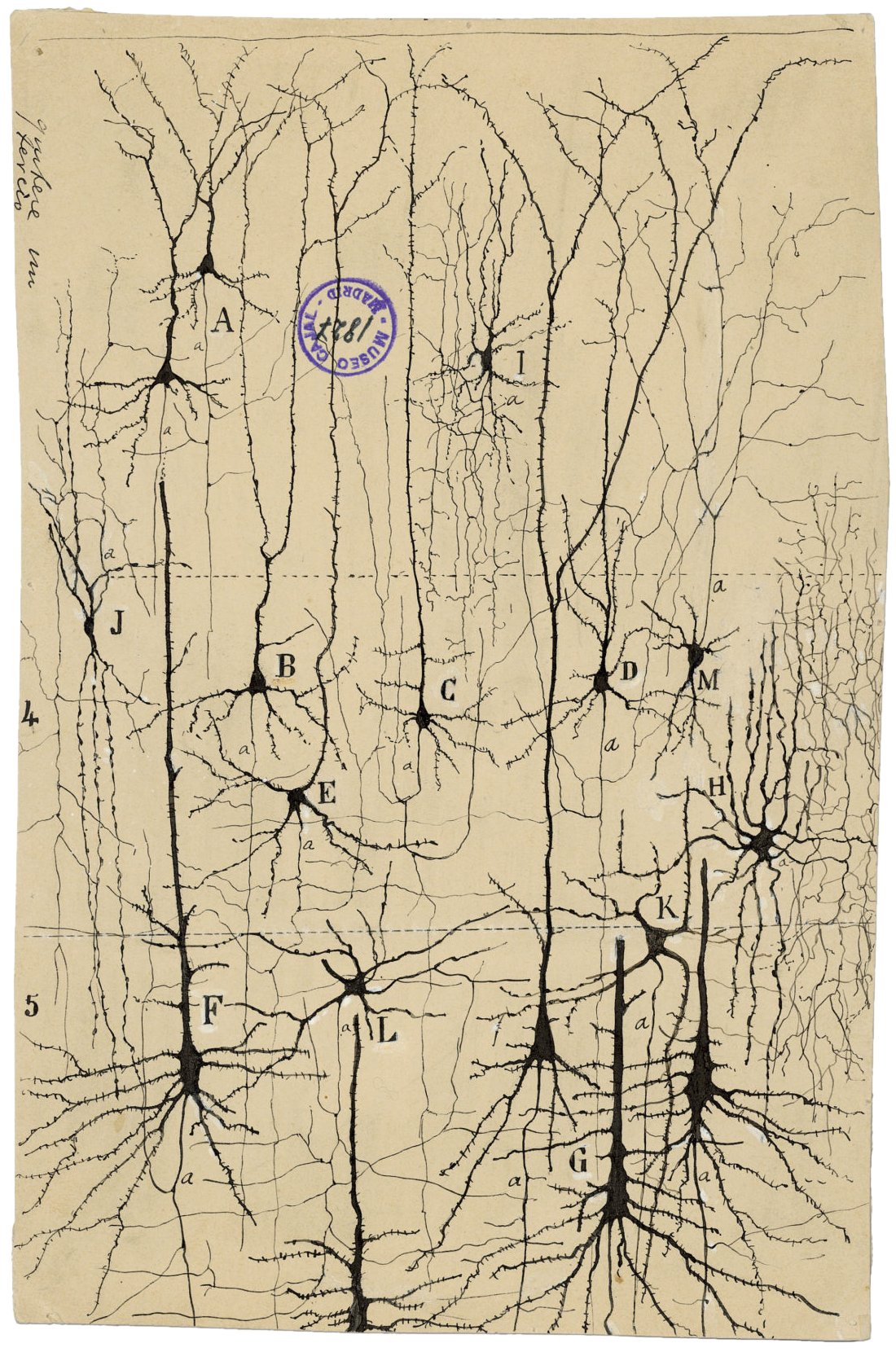
In the late 19th century, the Spanish pathologist Santiago Ramón y Cajal drew inky black lines to illustrate neurons in what can be considered the first brain cell atlas.
Courtesy of the Cajal Institute and the Spanish National Research Council
Some researchers have a larger goal in mind: They want to stitch all the maps together to create what Roser Vento-Tormo, the group leader of cellular genetics at the Wellcome Sanger Institute, compared to a “Google Maps” of the human body.
Like a biological version of Ortelius’ endeavor, the Human Cell Atlas project is working to draft such a map covering all 36 trillion cells in the human body by linking up reference maps contributed by scientists around the world. Founded in 2016, the project now claims more than 2,300 collaborators and high-profile funders such as the National Institutes of Health and the European Union, among many other scientific and private groups.
The first version of the human cell atlas is tantalizingly close to its big reveal: It should be done in around two years, Vento-Tormo said. It is unlikely to include profiles of all human cell types. Like the first geographic atlases, or the first published human genome, the first human cell atlas will be incomplete at its release, but with plans to revise it repeatedly. “We are at the beginning,” Vento-Tormo said.
Future cell atlases will probably also probe cell identity more deeply. Most cell maps are looking only at messenger RNA, which is “not even the only kind of RNA in the cell,” Quake said. Moreover, an expressed gene doesn’t necessarily mean that a specific protein is present in a cell. A future goal is to gather data on which proteins and other molecular products are present in single cells, Rhea said.
Even after the first full human cell map is done, the work won’t be finished. A central aim of the Human Cell Atlas project is to represent the full range of human diversity, which will require many samples from populations around the world to fully map our cellular diversity. Those samples will need to consider many axes of diversity: sex, age, ancestry, disease state and more.
The daunting work that lies ahead can’t quench researchers’ exhilaration over the current atlases. “It’s going to be very exciting to see what comes of all this over the next two or three years,” Angelo said. Already the first editions of these atlases are revealing how much we still have to learn about basic biology while also raising brand-new questions — clear signs of the beginning of a new scientific revolution.
“We will for sure learn tons from these efforts,” Naselaris said. “We just don’t know what we’ll learn because we lack the basic facts needed to even formulate most of the important questions.” Like geographers’ first atlases, where rivers, mountains and sometimes entire countries were missing, these atlases are early drafts of what’s to come.